Discovery increases chance of improving iron content in plants
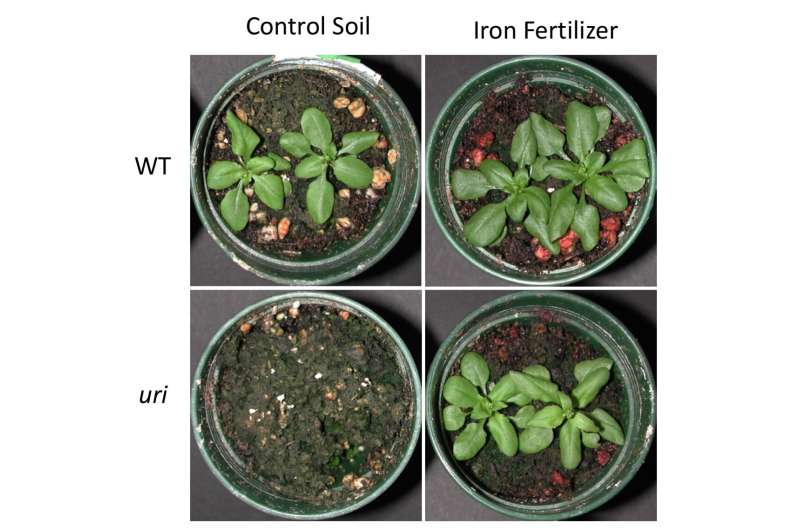

Researchers have discovered a gene that controls the regulation of iron uptake in plants, according to a new study from Dartmouth College.
With over 2 billion people suffering from iron deficiency around the world, the discovery could be the key to increasing the iron potency of crops such as rice, wheat and cassava that form the staple diets of more than half the world's population.
The gene signals when plants should allow the transport of iron by toggling a genetic pathway that includes hundreds of other genes.
The study, published in Proceedings of the National Academy of Sciences, is the latest in a decades-long series of research projects on iron transport and storage in plants from the Dartmouth team. The current research details the identification of the gene in plants and provides the first description of how it works to regulate iron uptake.
"We have discovered a key regulator in one of the world's most important nutrient pathways," said Mary Lou Guerinot, a professor of biological sciences at Dartmouth and the senior researcher on the project. "If we can now figure out how to optimize the pathway, we could feed billions of people that suffer from iron deficiency—a huge problem especially for women and children worldwide."
The newly-discovered gene, known as Upstream Regulator of IRT1 (URI), controls when genes should be expressed in the root of a plant to begin iron uptake. According to the team, URI controls as many as 1,500 other genes, including those that have nothing to do with iron.
The research team found that the URI protein is always present in plants. The constant presence of the protein created the additional challenge of understanding how the protein senses the status of iron sufficiency in order to regulate itself and avoid toxic overexposure to the element.
"Our research found that the abundance of the URI protein is not changed by iron conditions," said Sun A Kim, a research scientist at Dartmouth and the paper's first author. "Faced with the finding that the protein is always present, we went on to investigate if the URI protein is modified in response to iron availability to alter its activity."
Researchers found that under iron-poor conditions, the URI protein combines with a phosphate molecule and activates a sequence of genetic events to turn on the iron uptake system. This phosphorylation process is a common mechanism used by cells to regulate protein function and transmit signals.
URI also controls a protein that shuts down the transport of iron through the plant root under iron-sufficient conditions.
"Not only did we find URI, we found the regions of DNA where the URI protein binds to allow the iron-uptake pathway to be expressed. URI is a transcription factor and it's being modified to know when to bind and turn on other genes," said Kim.
Plants rely on iron for photosynthesis and growth. But iron is highly reactive and toxic, so uptake and storage of the element is tightly controlled. The result is that plants tend to overregulate iron.
The research team is now focusing on developing a plant with a regulator that stays in the "on" position for a longer amount of time. The team's goal is to have plants take up enough iron to benefit human consumers of plants.
"Foods in many of the staple diets around the world are not a good source of iron like red meat. This research discovery could provide iron for people who are not already getting it," said Guerinot.
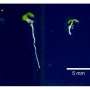
Past research by the team to identify mechanisms that control the transport and storage of iron in plants has resulted in crops that accumulate more iron. The current research was conducted on the model plant Arabidopsis thaliana and adds a crucial piece to the puzzle of how plants regulate iron uptake.
More information: Sun A. Kim el al., "The iron deficiency response in Arabidopsis thaliana requires the phosphorylated transcription factor URI," PNAS (2019). www.pnas.org/cgi/doi/10.1073/pnas.1916892116
Journal information: Proceedings of the National Academy of Sciences
Provided by Dartmouth College