Getting to the root of how plants tolerate too much iron
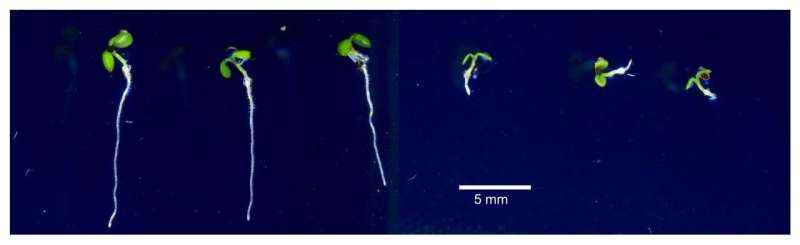
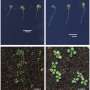
Iron is essential for plant growth, but with heavy rainfall and poor aeration, many acidic soils become toxic with excess iron. In countries with dramatic flood seasons, such as in West Africa and tropical Asia, toxic iron levels can have dire consequences on the availability of staple foods, such as rice.
Despite dozens of attempts in the last two decades to uncover the genes responsible for iron tolerance, these remained elusive until recently. Now, Salk scientists have found a major genetic regulator of iron tolerance, a gene called GSNOR. The findings, published in Nature Communications on August 29, 2019, could lead to the development of crop species that produce higher yields in soils with excess iron.
"This is the first time that a gene and its natural variants have been identified for iron tolerance," says Associate Professor Wolfgang Busch, senior author on the paper and a member of Salk's Plant Molecular and Cellular Biology Laboratory as well as its Integrative Biology Laboratory. "This work is exciting because we now understand how plants can grow in stressful conditions, such as high levels of iron, which could help us make more stress-resistant crops."
In plants such as rice, elevated soil iron levels cause direct cellular damage by harming fats and proteins, decreasing roots' ability to grow. Yet, some plants appear to have inherent tolerance to high iron levels; scientists wanted to understand why.

"We believed there were genetic mechanisms that underlie this resistance, but it was unclear which genes were responsible," says first author Baohai Li, a postdoctoral fellow in the Busch lab. "To examine this question, we used the power of natural variation of hundreds of different strains of plants to study genetic adaption to high levels of iron."
The scientists first tested a number of strains of a small mustard plant (Arabidopsis thaliana), to observe if there was natural variation in iron resistance. Some of the plants did exhibit tolerance to iron toxicity, so the researchers used an approach called genome-wide association studies (GWAS) to locate the responsible gene. Their analyses pinpointed the gene GSNOR as the key to enabling plants and roots to grow in iron-heavy environments.
The researchers also found that the iron-tolerance mechanism is, to their surprise, related to the activities of nitric oxide, a gaseous molecule with a variety of roles in plants including responding to stress. High levels of nitric oxide induced cellular stress and impaired the plant roots' tolerance for elevated iron levels. This occurred when plants did not have a functional GSNOR gene. GSNOR likely plays a central role in nitric oxide metabolism and regulates the plants' ability to respond to cellular stress and damage. This nitric oxide mechanism and the GSNOR gene also affected iron tolerance in other species of plants, such as rice (Oryza sativa) and a legume (Lotus japonicus), suggesting that this gene and its activities are likely critical in many, if not all, species of plants.
"By identifying this gene and its genetic variants that confer iron tolerance, we hope to help plants, such as rice, become more resistant to iron in regions with toxic iron levels," says Busch. "Since we found that this gene and pathway was conserved in multiple species of plants, we suspect they may be important for iron resistance in all higher plants. Additionally, this gene and pathway may also play a role in humans, and could lead to new treatments for conditions associated with iron overload."
Next, Li will be starting his own laboratory at Zhejiang University, in China. He plans to identify the relevant genetic variants in rice and observe if iron-tolerance variants could increase crop yields in flooded Chinese fields.
More information: Baohai Li et al, GSNOR provides plant tolerance to iron toxicity via preventing iron-dependent nitrosative and oxidative cytotoxicity, Nature Communications (2019). DOI: 10.1038/s41467-019-11892-5
Journal information: Nature Communications
Provided by Salk Institute