Sewage, rivers and soils provide missing link in antibiotic resistance story

If you think that the key to beating antibiotic resistance is only for doctors to prescribe less and scientists to find new drug candidates, you are probably wrong. The fundamental solutions may lie far from medicine—in managing our rivers and soils.
That is the view of scientists who have uncovered disturbing amounts of antibiotics, and antibiotic-resistant bacteria, in rivers and wastewater over the last decade. Now they are developing techniques to quantify the dangers contained within any particular body of water—and they hope this may provoke policymakers into action.
One way in which antibiotic residues reach the environment is when people excrete them in their faeces and urine, says Professor Willem van Schaik, a specialist in microbiology and infection at the University of Birmingham in the UK. And it's not just the drugs: the germs causing the disease the antibiotics were taken for will hitch a ride too—and a portion of these will be resistant to the drugs.
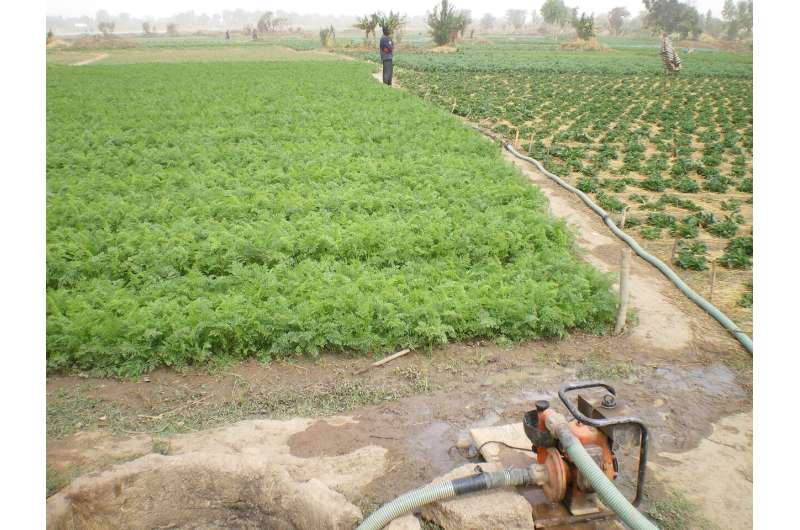
In poorer countries, more than 80% of sewage is discharged untreated into the environment. In Cameroon's capital, Yaounde, Dr. Blaise Bougnom, an environmental microbiologist at the University of Yaounde, has been piecing together one particular route for the sewage: human waste often goes directly into city canals, from where urban farmers take water to irrigate crops crucial for city-dwellers' vegetable supplies.
Urban farming
Urban farming is widespread in poorer countries, with vegetables growing on roadsides, along drainage canals, in roundabouts and parks. It can provide 40% of a city's food supply and up to 90% of its demand for vegetables.
Dr. Bougnom says there is an 'alarming' level of antibiotic-resistant infections in Yaounde's hospitals.
"I discussed with some medical doctors," he said. "I started thinking that there might be a relationship between what we are seeing in the clinical setting and what is happening in the environment. Then I developed this project looking at the potential of wastewater to transmit AMR (antimicrobial resistance)."
He began the research while at the University of Birmingham studying canal wastewater used for irrigation in the cities of Ouagadougou, Burkina Faso, and Yaounde and Ngaoundere in Cameroon. He analysed which genes were present in the water and found many related to bacteria that cause diarrhoea in humans. He also found many genes related to antibiotic resistance, including more than 80 that are known to be easily transferred between bacteria, along with entities, such as plasmids, that transfer them.
He then compared urban fields in these cities, finding that those that had been irrigated with raw wastewater held more antibiotics and resistance genes than those that had not.
"It's very tragic because something that was supposed to bring many solutions (urban farming), in fact also might bring many problems," he said.
Another potent source of resistance-laden wastewater may be hospital sewage.
Across the world, antibiotic use is likely to be higher in hospitals—in Europe, a third of hospital patients are taking antibiotics at any one time, according to Prof. van Schaik, while the figure for the United States is a half. Thus, the load in the wastewater—both of antibiotics and resistant bacteria—is likely to be high.
"You can think of it (the hospital sewerage system) as a pipe where an entire bacterial ecosystem is growing on the walls," said Prof. van Schaik. "These ecosystems will be optimised to grow in the conditions of the sewage that's flowing through those systems and because levels of antibiotics will be very high—almost higher than anywhere else—those bacteria will have adapted to life in the presence of high levels of antibiotics."
Just how loaded the wastewater is—and therefore what size of a problem we face—is unknown, but Dr. Lisandra Zepeda Mendoza, a genomics expert and colleague of Prof. van Schaik, is hoping to change that by studying the genetic material of the microbes in that ecosystem in a project called ARCS.
Risky
She has been analysing the DNA from the assorted bacteria and viruses found in jars of hospital wastewater, looking for elements such as genes associated with the transfer of resistance between bacteria and phages—viruses that can transfer genetic material between bacteria. To understand how risky a sample is, it's important to understand not just which resistance genes are present but also how transmissible they are, which depends on what else is present.
"I can identify what bacteria are there and what genes they have and what they can do," she said.
She is feeding these results—as well as data from other analyses of hospital wastewater—into a machine-learning model that can deduce how potent the resistance threat is in any particular source.
If her approach is successful, it could be used to compare the relative dangers of different sources—for example, rivers close to drug factories or hospitals, city canals and irrigation water.
"There's not really a way to compare different environments in a standardised way," she said.
Prof van Schaik added: "People in my field are having massive arguments about what are the most effective interventions and what are the most important things to be working on and the short answer is we don't know.
"I think with this approach we will at least be able to make a risk assessment of different environments—and not just wastewater but the bacteria that colonise the human gut or the animal gut or the soil or anything you can think of.
"This could settle arguments about the differing contributions a variety of practices make to generating antibiotic resistance.
"If you go to lower- and middle-income countries, antibiotic resistance really is a massive clinical issue and also very costly. If we can maybe with this study at least convince some people that it is a good investment to try and address the underlying cause of antibiotic resistance, rather than (just) trying to improve treatments for patients in hospitals, that would be a very good policy outcome."
Provided by Horizon: The EU Research & Innovation Magazine