Team's study reveals details of new DNA repair pathway

A team of Vanderbilt investigators has discovered how a DNA repair pathway protein shields sites of damage to avoid mutations and maintain genome integrity.
"DNA repair is critical to prevent cancer. The loss of or inefficiencies in DNA repair cause mutations and changes in our chromosomes that lead to cancer development," said David Cortez, Ph.D., Ingram Professor of Cancer Research and professor of Biochemistry.
Earlier this year, Cortez and his colleagues reported a new DNA repair mechanism, involving a protein called HMCES. Now, in collaboration with Brandt Eichman, Ph.D., and his group, the researchers have uncovered structural details of how HMCES binds to damaged DNA to protect it from mutation. Their findings were reported in the journal Nature Structural & Molecular Biology.
"Not only is this a cool scientific story, it was also a beautiful collaboration between two graduate students," said Eichman, William R. Kenan, Jr. Professor and chair of the Department of Biological Sciences.
The Cortez team previously showed that HMCES has a role in the repair of "abasic sites"—the most common type of DNA damage—in single-stranded DNA. Abasic sites happen as many as 20,000 times per day in human cells. When HMCES is missing, cells accumulate DNA damage and have increased genetic stability.
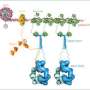
In studies led by postdoctoral fellow Kareem Mohni, Ph.D., the researchers demonstrated that HMCES binds to an abasic site and forms a DNA-protein cross-link.
Petria Thompson, a graduate student in the Cortez laboratory, was interested in understanding more about the chemical nature of the cross-link between HMCES and an abasic site. Using biochemical experiments, she showed that the cross-link was extremely stable, and she developed methods for purifying large quantities of the DNA-protein cross-link.
Thompson then teamed with Katherine Amidon in the Eichman laboratory for structural studies.
The pair purified DNA-protein cross-links using two proteins: human HMCES and a homologous protein from E. coli. Amidon produced high quality crystals of both proteins bound to DNA and used X-ray crystallography to determine a high-resolution structure of the DNA-protein cross-link.
The type of chemical bonds they identified between the protein and the abasic site explained the remarkable stability of the DNA-protein cross-link.
Thompson and Amidon "worked on all aspects of protein purification, crystallization and biochemical experiments by sharing their expertise, data and reagents," Eichman said. "This collaboration enabled the project to progress much more efficiently than it would have otherwise."
The structure revealed that HMCES has specificity for abasic sites in single-stranded DNA at junctions that occur during replication—when a polymerase protein that is copying the DNA encounters an abasic site. The findings support a role for HMCES in protecting abasic sites during DNA replication.
"From our earlier biochemical studies, we predicted that HMCES would have a binding preference for these types of abasic sites," Cortez said. "It's exciting that the structural biology studies have confirmed our predictions.
How repair of the abasic site proceeds following formation of the DNA-protein cross-link is an open question and the subject of future studies.
More information: Petria S. Thompson et al. Protection of abasic sites during DNA replication by a stable thiazolidine protein-DNA cross-link, Nature Structural & Molecular Biology (2019). DOI: 10.1038/s41594-019-0255-5
Journal information: Nature Structural & Molecular Biology
Provided by Vanderbilt University