Finding missing network links could help develop new drugs, stop disease, ease traffic
A new mathematical model of the structure of networks could help find new cancer drugs, speed up traffic flow and combat sexually transmitted disease.
Although the three challenges seem diverse, they all could benefit from a theory that helps to uncover unknown information about a network by analyzing its structure. The study was published in the journal Chaos.
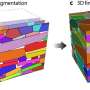
An example of how filling in missing links could be useful is in choosing a genetic target for a cancer drug, said team member Michael Small, from the University of Western Australia.
"Say you have a network of genes that are somehow connected, and there are some known drug targets. But if you don't know all the genes, you would like to make guesses at information that you don't have to work out what might be other likely targets to investigate," Small said.
Successful link prediction algorithms already exist for certain types of networks, but the researchers analyzed differently structured networks to come up with their alternative algorithm.
An example of traditional link prediction might be a social network suggesting friends with which to connect. Two people who share a friend are likely to themselves be friends with a direct connection. Adding this third connection creates a triangle, which is a structure where existing algorithms focus.
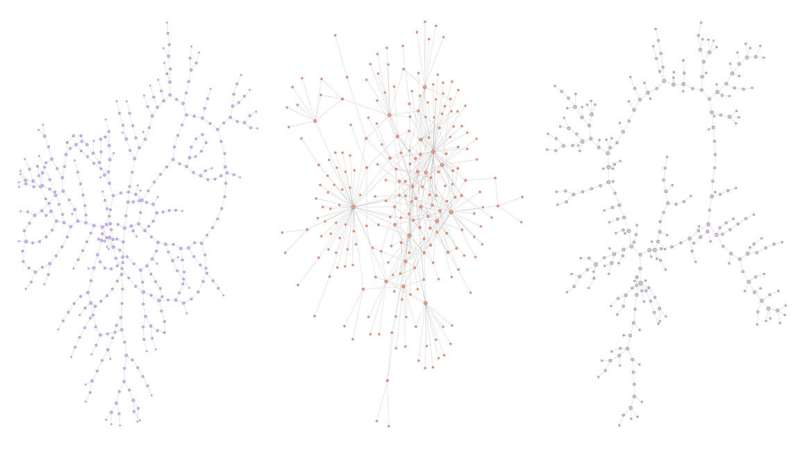
In contrast, the authors focused on treelike networks, which have many branches but very few cross links between branches. They studied three example datasets: the social network Twitter, a water distribution network and a sexual contact network.
They found these three treelike networks could be characterized by a number of parameters, such as the average distance between branch points in the network, the size of loops and a comparison of the number of links that adjacent nodes have—a measure of the network's regularity describing heterogeneity.
The authors then developed an algorithm that suggested links that would preserve these characteristics of the network.
They tested this approach by taking a known network and removing links from it and seeing if the algorithm could predict where the missing links should be.
The team found the algorithm performed better (around 44% for the Twitter network, around 15% for the sexual contact network and around 4% for the water distribution network) for treelike networks than most of conventional algorithms which rely on other parameters, such as assuming highly connected individuals will attract more connections (known as preferential attachment) or building lots of triangular connections (known as clustering).
Author Keke Shang attributes the team's success to thinking about examples of real-world networks.
"I hope we can make network technology serve our lives better," he said.
More information: "Link prediction for tree-like networks," Chaos (2019). DOI: 10.1063/1.5107440
Journal information: Chaos
Provided by American Institute of Physics