Researchers develop fast, efficient way to build amino acid chains

Scientists often build new protein molecules by stringing groups of amino acids together. These amino acid chains, called polypeptides, are the building blocks needed in drug development and the creation of new biomaterials.
The process for building polypeptides is difficult, however. Researchers report that they have developed a faster, easier and cheaper method for making new polypeptides than was previously available. The new approach uses a streamlined process that purifies the amino acid precursors and builds the polypeptides at the same time, unlike previous methods in which the processes were separate, laborious and time-consuming.
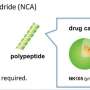
Traditionally, making polypeptide chains has been a very complicated process, said University of Illinois materials science and engineering professor Jianjun Cheng, who led the new research. Synthesizing and purifying the amino acid precursors, namely N-carboxyanhydride, or NCA, requires days of tedious effort, and building the polypeptide chains takes hours to days, he said.
"The field has never grown big, in part because synthesizing polypeptides is so complicated," Cheng said. "NCA has a lot of impurities that are difficult to remove. Until now, the synthesis of high-quality polypeptides required ultrapure NCAs."
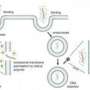
In biological cells, enzymes called ribozymes join amino acids together to form proteins, Cheng said. This process takes place in the presence of water, salt and numerous other molecules. Replicating this process in the laboratory is very difficult, however. Current methods require researchers to use purified NCA molecules and to build the chains in a water-free environment.
Cheng and his colleagues drew inspiration from ribozymes, which excel at making amino acid chains quickly while isolating them from the cellular environment. The team developed a system that mimics the ribozyme function, building the amino acid chains quickly while removing any molecules that could contaminate the system. This allows the researchers to build the desired chains with NCAs that are not pure.
"This is the first time since the discovery of the NCA molecule in 1906 that we have been able to build long chains using non-purified NCA," Cheng said.
"I worked on NCA purification for several years and found it very painful, because the process required water-free conditions and was technically challenging," said postdoctoral researcher Ziyuan Song, a member of Cheng's lab. "That's why there aren't many research groups working in this field. With this method, we can get more people to join and find more applications."
The method can be used in chemistry, biology and industry, where protein chains are routinely used as building blocks for the assembly of useful molecules, the researcher said.
"Previously, the field required specialized chemists like us to make these building blocks," Cheng said. "Our new protocol allows anyone with basic chemistry skills to build the desired polypeptides in a few hours."
The researchers are investigating how to scale up the process and explore the full range of chemical and biological applications the new approach allows.
The researchers report their findings in the Proceedings of the National Academy of Sciences.
More information: Ziyuan Song et al, Synthesis of polypeptides via bioinspired polymerization of in situ purified N-carboxyanhydrides, Proceedings of the National Academy of Sciences (2019). DOI: 10.1073/pnas.1901442116
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Illinois at Urbana-Champaign