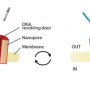
Major advance in nanopore detection of peptides and proteins
Nanopore technology, which is used to sequence DNA, is cheap, hand-held and works in the jungle and in space. The use of this technology to identify peptides or proteins is now a step closer. University of Groningen scientists have used a patented nanopore to identify the fingerprints of proteins and peptides, and it can even detect polypeptides differing by one amino acid. The results were published on 16 October in the journal Nature Communications.
University of Groningen scientists have been able to identify a number of peptides and proteins passing through a funnel-shaped nanopore. They have solved two main problems that have hampered attempts to analyze and sequence proteins with nanopores: getting polypeptides into the pore and identifying differences in proteins by recordings of current. 'Nanopores usually carry a charge, and the amino acids that make up polypeptides are also charged. Getting the polypeptide inside the pore and to pass through nanopores is therefore a challenge', explains associate professor of Chemical Biology Giovanni Maglia.
Fingerprint
He used an electro-osmotic flow to pull the polypeptides into the pores. Under an applied potential across the nanopore, a flow of ions and water passes through the pore.' If the direction of the ion current can be controlled, a fluid flow strong enough to transport polypeptides can be generated. 'We did this by tuning the charges inside the pore wall. By changing the pH of the medium, it was possible to fine-tune the balance between the electro-osmotic flow and the force of the electric field which was applied across the pore.'
Maglia tested five different polypeptides ranging from 1 to 25 kilodalton. 'We used biomarker peptides linked to disease, with different charges and shapes', he says. The polypeptides entered the pore and the current across the pore produced a 'fingerprint' for each. He thus managed to distinguish two versions of the 21 amino acid peptide endothelin, which differ by just one amino acid (tryptophan or methionine).
Sequencing
Getting a good reading from a nanopore is complicated. Maglia used a new kind of pore that he characterized and patented. 'Pores used in the past are barrel-shaped, which means the shape and size of the pore has fundamental limitations. But our pore has an alpha helical funnel shape, and the size of the narrow end, which is where we do our measurements, means it should contain just one amino acid, so it is more easily tuned.'
Currently, the polypeptides pass through the pore too rapidly to identify the separate amino acids. This is needed for protein sequencing at the single-molecule scale. It would be a valuable tool for research, explains Maglia: 'Proteins can be chemically modified in many unique ways, and we have very little information on the exact composition of proteins in our body.' This can only be seen at the single-molecule level.
Maglia: 'Molecular diagnostics and biomarker discovery should benefit particularly from the single-molecule characterization of proteomes.' It is a major advantage that nanopore technology has already been developed for DNA sequencing. This technology is fast, cheap and robust: nanopore sequencing devices are used in the field and one has even been sent up to the International Space Station. Using a similar technique to identify proteins would require minor adaptations, mainly in the pores. 'In theory, we could build an application tomorrow.'
More information: Gang Huang et al, Electro-osmotic capture and ionic discrimination of peptide and protein biomarkers with FraC nanopores, Nature Communications (2017). DOI: 10.1038/s41467-017-01006-4
Journal information: Nature Communications
Provided by University of Groningen