Creating a global map of the protein shape universe
Proteins can provide a detailed look inside the human body and how it protects itself from many diseases. Proteins, which make up about 15% of body mass, are the most abundant solid substances in the human body. They are important working molecules of the immune system, metabolism, brain function, body motion, and any physically and chemically functional parts in a body. Each protein has a specific function under the direction of its own gene.
Now, Purdue University researchers have come up with a novel way to classify proteins and their shapes, which lays the foundation of how we understand protein structures and functions. The shapes are important because they determine the role and effectiveness of the proteins. The research is published in the April edition of PLOS Computational Biology.
"We developed a new way to view and classify protein 3-D shapes, which provides our basic understanding of protein shapes and will become a foundation of artificial protein design and other applications of proteins," said Daisuke Kihara, a professor of biological sciences and computer science in Purdue's College of Science, who leads the research team.
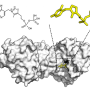
Kihara said proteins were conventionally classified by conformations of protein chains, a rather detailed level of structural features. They currently classify proteins by their overall surface shapes, which would be more directly relevant to how they interact with other proteins and compounds in a cell.
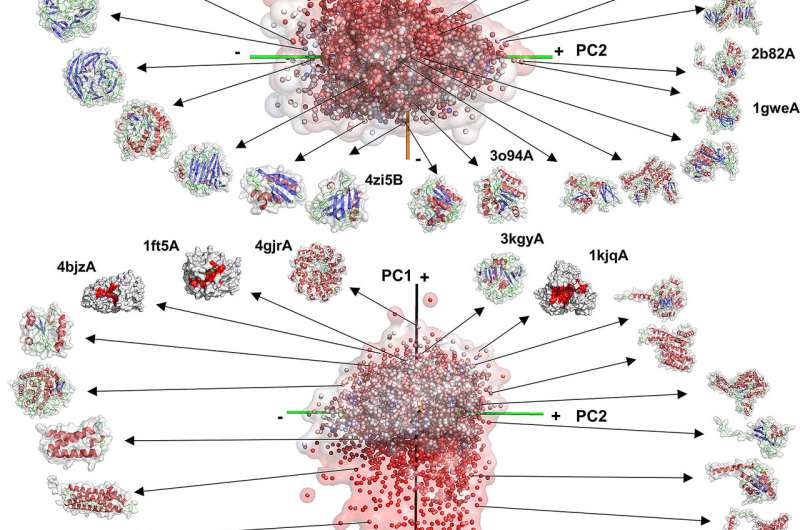
The Purdue team maps and classifies 3-D surface shapes of proteins within a certain space inside the body. Surface shapes of proteins are represented with 3-D Zernike descriptors, mathematical moment-based invariants, which have previously been demonstrated effective for biomolecular structure similarity search.
They also have analyzed the shape space occupied by protein complexes. The mapping provides data into the relationship between shapes, main-chain folds and complex formation.
"This work is in the area of bioinformatics," Kihara said. "Bioinformatics is not only useful for processing biological data or developing computational tools, but it is also helpful for providing a unique view and framework that can change the way people view and understand the biomolecular world."
Kihara has worked with the Purdue Research Foundation Office of Technology Commercialization on some of his research and technology.
His team's work aligns with Purdue's Giant Leaps celebration, celebrating the global advancements in health research and technology as part of Purdue's 150th anniversary. Health is one of the four themes of the yearlong celebration's Ideas Festival, designed to showcase Purdue as an intellectual center solving real-world issues.
More information: Xusi Han et al, A global map of the protein shape universe, PLOS Computational Biology (2019). DOI: 10.1371/journal.pcbi.1006969
Journal information: PLoS Computational Biology
Provided by Purdue University