Key differences between prokaryotic and eukaryotic RNA silencing Argonaute enzyme unveiled
The Argonaute (Ago) enzyme complex plays a critical role in DNA and RNA target cleavage for a process known as RNA silencing in prokaryotic and eukaryotic cells, making them a target for future gene-editing technology. The present study unravels key differences between prokaryotic Ago (pAgo) and eukaryotic Ago (eAgo) enzymes in the cleavage reaction and may provide important clues on their evolutionary past.
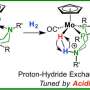
Enzymes have clearly defined active sites to allow the substrate molecule to fit intricately. This is often coupled with an enzymatic conformational change prior to the occurrence of the catalysis reaction. For Ago, the catalysis step requires insertion of a "glutamate finger" to form the catalytic plugged-in conformation, which can be stabilized through hydrogen-bonding networks provided by two symmetrical, positively-charged residues.
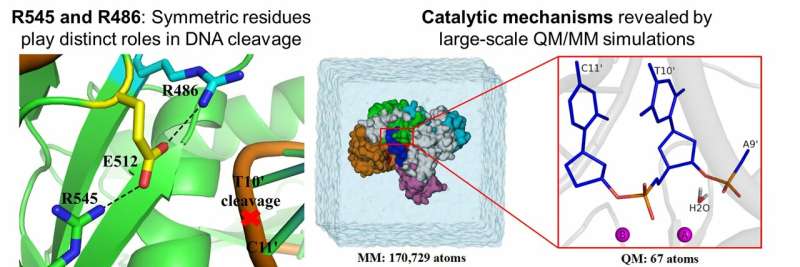
For Ago in eukaryotes, these two symmetric positively-charged residues play an identical role that is critical for cleavage. Hence, researchers have speculated that the two analogous resides in prokaryotic Ago perform the same critical role in cleavage function. Surprisingly, this study (Fig. 1) showed that in pAgo, only one (Arginine 545) of the two residues is involved in cleavage function. When the other one (Arginine 486) was substituted with other amino acids, the enzyme was still able to maintain its cleavage activity. Based on these results, the study further suggested that R486 may play other roles such as assisting the insertion of the glutamate finger. The discovery of such striking differences in the roles of these symmetric resides between eAgos and pAgos provides novel insights on how the cleavage functions evolve during the evolution journey from prokaryote to eukaryote.
To achieve these results, computational methods combining quantum mechanics, molecular mechanics, and molecular dynamics (QM/MM) were applied to elucidate the cleavage reaction mechanism and identify functional roles of the amino acid residues. This research was made possible by large-scale, high-performance computing resources, which were computed equivalent to 10,000 CPU cores for 25 weeks on the Shaheen II Supercomputer at KAUST in collaboration with Prof. Xin Gao's group.
"This research was made possible due to current day computing capabilities and the precision that QM/MM modelling allows for," said Prof. Huang Xuhui. "Comparing which amino acid residues play a key part in the target DNA/RNA cleavage step in pAgo and eAgo sheds light on how Ago protein evolves from prokaryotes to eukaryotes to cleave DNA/RNA. This information may be useful in ultimately modifying the Ago protein for use as an enhanced gene editing tool in the future," Prof. Huang explained.
Details of the methodology and their findings were published in the Proceedings of the National Academy of Sciences (PNAS) journal on December 27, 2018.
More information: Jinping Lei et al, Two symmetric arginine residues play distinct roles in Thermus thermophilus Argonaute DNA guide strand-mediated DNA target cleavage, Proceedings of the National Academy of Sciences (2018). DOI: 10.1073/pnas.1817041116
Journal information: Proceedings of the National Academy of Sciences
Provided by Hong Kong University of Science and Technology