Detangling DNA replication
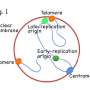
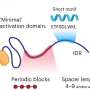
DNA is a lengthy molecule—approximately 1,000-fold longer than the cell in which it resides—so it can't be jammed in haphazardly. Rather, it must be neatly organized so proteins involved in critical processes can access the information contained in its nucleotide bases. Think of the double helix like a pair of shoe laces twisted together, coiled upon themselves again and again to make the molecule even more compact.
However, when it comes time for cell division, this supercoiled nature makes it difficult for proteins involved in DNA replication to access the strands, separate them, and copy them so one DNA molecule can become two.
Replication begins at specific regions of the chromosome where specialized proteins separate the two strands, pulling apart the double helix as you would the two shoe laces. However, this local separation actually tangles the rest of the molecule further, and without intervention creates a buildup of tension, stalling replication. Enter the enzymes known as topoisomerases, which travel ahead of the strands as they are being peeled apart, snipping them, untwisting them, and then rejoining them to relieve the tension that arises from supercoiling.
These topoisomerases are generally thought to be sufficient to allow replication to proceed. However, a team of researchers from MIT and the Duke University School of Medicine suggests the enzymes may require guidance from additional proteins, which recognize the shape characteristic of overtwisted DNA.
"We've known for a long time that topoisomerases are necessary for replication, but it's never been clear if they were sufficient on their own," says Michael Laub, an MIT professor of biology, Howard Hughes Medical Institute Investigator, and senior author of the study. "This is the first paper to identify a protein in bacteria, or eukaryotes, that is required to localize topoisomerases ahead of replication forks and to help them do what they need to do there."
Postdoc Monica Guo '07 and former graduate student Diane Haakonsen Ph.D. '16 are co-first authors of the study, which appeared online in the journal Cell on Sept. 13.
Necessary but not sufficient
Although it's well established that topoisomerases are crucial to DNA replication, it is now becoming clear that we know relatively little about the mechanisms regulating their activity, including where and when they act to relieve supercoiling.
These enzymes fall into two groups, type I and type II, depending on how many strands of DNA they cut. The researchers focused on type II topoisomerases found in a common species of freshwater bacteria, Caulobacter crescentus. Type II topoisomerases in bacteria are of particular interest because a number of antibiotics target them in order to prevent DNA replication, treating a wide variety of microbial infections, including tuberculosis. Without topoisomerases, the bacteria can't grow. Since these bacterial enzymes are unique, poisons directed at them won't harm human topoisomerases.
For a long time, type II topoisomerases were generally assumed adequate on their own to manage the overtwisted supercoils that arise during replication. Although researchers working in E. coli and other, higher organisms have pinpointed additional proteins that can activate or repress these enzymes, none of these proteins were required for replication.
Such findings hinted that there might be similar interactions taking place in other organisms. In order to understand the protein factors involved in compacting Caulobacter DNA—regulating topoisomerase activity specifically—the researchers screened their bacteria for proteins that bound tightly to supercoiled DNA. From there, they honed in on one protein, GapR, which they observed was essential for DNA replication. In bacteria missing GapR, the DNA became overtwisted, replication slowed, and the bacteria eventually died.
Surprisingly, the researchers found that GapR recognized the structure of overtwisted DNA rather than specific nucleotide sequences.
"The vast majority of DNA-binding proteins localize to specific locations of the genome by recognizing a specific set of bases," Laub says. "But GapR basically pays no attention to the actual underlying sequence—just the shape of overtwisted DNA, which uniquely arises in front of replication forks and transcription machinery."
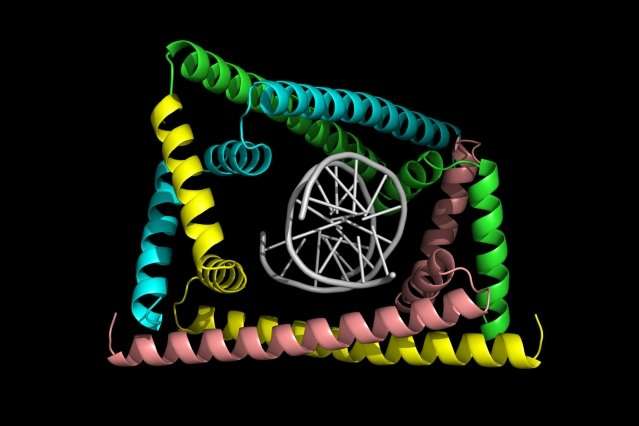
The crystal structure of the protein bound to DNA, solved by Duke's Maria Schumacher, revealed that GapR recognizes the backbone of DNA (rather than the bases), forming a snug clamp that encircles the overtwisted DNA. However, when the DNA is relaxed in its standard form, it no longer fits inside the clamp. This might signify that GapR sits on DNA only at positions where topoisomerase is needed.
An exciting milestone
Although GapR appears to be required for DNA replication, it's still not clear precisely how this protein promotes topoisomerase function to relieve supercoiling.
"In the absence of any other proteins, GapR is able to help type II topoisomerases remove positive supercoils faster, but we still don't quite know how," Guo says. "One idea is that GapR interacts with topoisomerases, recognizing the overtwisted DNA and recruiting the topoisomerases. Another possibility is that GapR is essentially grabbing onto the DNA and limiting the movement of the positive supercoils, so topoisomerases can target and eliminate them more quickly."
Anthony Maxwell, a professor of biological chemistry at the John Innes Centre who was not involved with the study, says the buildup of DNA supercoils is a key problem in both bacterial replication and transcription.
"Identifying GapR and its potential role in controlling supercoiling in vivo is an exciting milestone in understanding the control of DNA topology in bacteria," he says. "Further work will be required to show how exactly these proteins cooperate to maintain bacterial genomic integrity."
According to Guo, the study provides insight into a fundamental process—DNA replication—and the ways topoisomerases are regulated, which could extend to eukaryotes.
"This was the first demonstration that a topoisomerase activator is required for DNA replication," she says. "Although there's no GapR homolog in higher organisms, there could be similar proteins that recognize the shape of the DNA and aid or position topoisomerases."
This could open up a new field of drug research, she says, targeting activators like GapR to increase the efficacy of existing topoisomerase poisons to treat conditions like respiratory and urinary tract infections. After all, many topoisomerase inhibitors have become less effective due to antibiotic resistance. But only time will tell; there is still much to learn in order to untangle the complex process of DNA replication, along with its many twists and turns.
More information: Monica S. Guo et al. A Bacterial Chromosome Structuring Protein Binds Overtwisted DNA to Stimulate Type II Topoisomerases and Enable DNA Replication, Cell (2018). DOI: 10.1016/j.cell.2018.08.029
Journal information: Cell
Provided by Massachusetts Institute of Technology
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.