Supercoil me! The art of knotted DNA maintenance
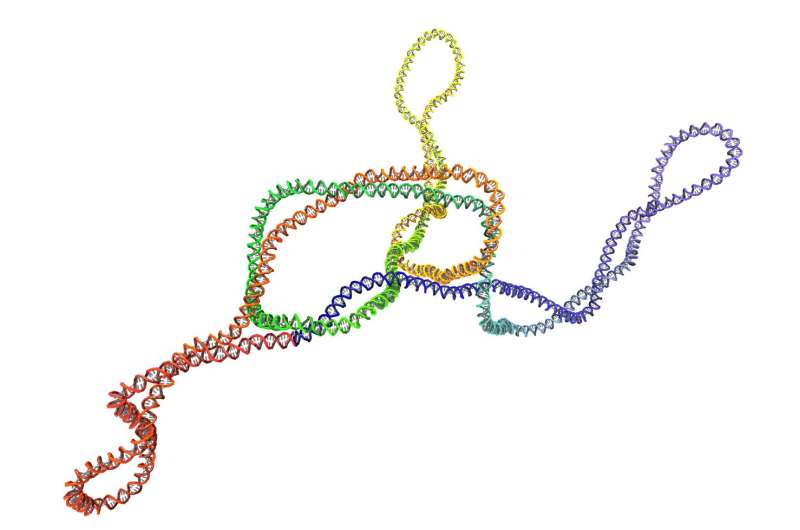
Even living cells have their own tangled problems to solve. They involve knots that arise accidentally in DNA and that can comprise its functionality. Researchers now suggest that the DNA propensity to be supercoiled, just like telephone cords, prevents accidental knots. These coils can keep DNA knots locked in place long enough to be untied by specialised enzymes. The computational study, based on molecular dynamics simulations of bacterial DNA, has just been published in Nucleic Acids Research.
A tangled problem
Supercoiled, twisted, and even knotted—far from the elegant and polished textbook images, DNA filaments in living cells is all crumpled and entangled, and to be functional, must be constantly unraveled, much like telephone cords that accumulate annoying curls or supercoils with careless use. Supercoils are present in DNA, too, and they are believed to be instrumental for keeping DNA compact and exposing or hiding genetic information to the cellular machinery that can process it, according to Lucia Coronel and Antonio Suma, young scientists and first authors of this study, which was coordinated by Cristian Micheletti.
At the same time, it has long been known that knots can accidentally form in DNA, with negative consequences for the cell. What had remained elusive until now was the dynamic interplay or coexistence on the same DNA filaments of complex knots and supercoiled regions. And this is what the researchers at SISSA set out to study using computer simulations, investigating the implications for DNA structure and its change in time, and whether such interplay might unlock new functional possibilities. They also sought to understand how the ensuing complex behaviour fits with what is already known about the molecular maintenance of DNA.
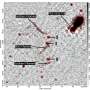
For this study, the scientists used detailed and extensive molecular dynamics simulations. They first noticed that in supercoiled DNA rings, knots are likely found in one of two positions—at the tip of the supercoils or in a more central position. A closer inspection revealed a more surprising and unexpected effect. Coronel and Suma write, "We studied the DNA filament and noted that without supercoils, the knotted region would move relatively fast along the filament. Likewise, supercoiled regions can rapidly change in knot-free DNA. However, when knots and supercoiling are simultaneously present, then the crucial contact points in DNA knots become locked in place, persistently. And this unexpected effect is particularly interesting because it could be key to a specific and unexpected biological functionality."
The authors write, "We know that living cells routinely deal with DNA knots, and we also know that these forms of entanglement are usually detrimental to biological functionality; for instance they can prevent the genetic information from being read and translated into protein products. Specific enzymes of the topoisomerase family are responsible for disentangling DNA. Their modus operandi recalls the efficient, if not drastic, way in which Alexander the Great untied the proverbial Gordian knot with a cut of his sword. Similarly, these enzymes untie DNA filaments by a sophisticated cutting and sealing action."
There is still an ongoing debate on which molecular mechanisms can guide these enzymes, which are minute compared to the typical scale of DNA filaments, to intervene on the correct places where their cut-and-paste action can untie the knots. The authors conclude, "It has been suggested that the target places are recognised by specific geometric features, which is a plausible and elegant mechanism. However, it was unclear how these features could persist in spite of the incessant molecular motion. Our study suggests that DNA supercoiling can favour the action of topoisomerases by keeping knots in a stable configuration for a time span that is much longer than other molecular rearrangements. In this way, the enzymes could have sufficient time to recognise the target sites and, in turn, their cut-and-paste action would be simpler, more reliable and efficient. This is presently a hypothesis but, since it has such interesting implications, that we hope can it be addressed in future experiments."
More information: Nucleic Acids Research (2018). DOI: 10.1093/nar/gky523/5042023
Journal information: Nucleic Acids Research
Provided by International School of Advanced Studies (SISSA)