Ants surrender their venomous secrets
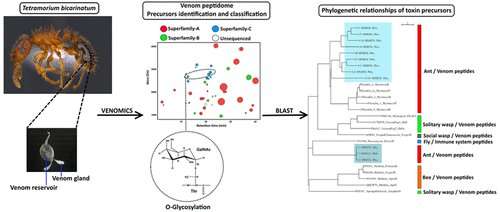
Venoms produced by snails, snakes, scorpions and spiders contain numerous bioactive compounds that could lead to therapeutic drugs or insect-specific pesticides. Yet little is known about venoms produced by insects, in part because each bug contains such a tiny amount. Researchers recently responded to this challenge by conducting one of the first intensive studies of ant venom. They have now published their findings in ACS' Journal of Proteome Research.
Ants produce venom for many reasons, such as defending themselves from predators, capturing prey or thwarting microbial infection. Prior studies have shown that venom is a complex cocktail that contains thousands of peptides, but fewer than 100 have been characterized in a couple of ant venoms. So Axel Touchard and colleagues set out to identify the peptides produced by a species of stinging ant known as Tetramorium bicarinatum, which is common in tropic and subtropical areas.
The researchers collected the material for their "venomics" study by dissecting ant venom sacs and glands. Using a wide range of techniques, they identified more than 2,800 venom peptides, most of which were fragments of 37 full-length peptide precursors called myrmicitoxins. The scientists sorted these precursors into three broad categories based on amino acid sequence; one category consists of an entirely new sequence never before observed in any venom. The researchers also showed that the myrmicitoxins shared some sequence similarities with peptides produced by other species, including other types of ants, as well as wasps, mosquitoes and fruit flies. The team says that venom peptides likely evolved from a relatively small set of ancestral genes that have been impacted by exposure to different prey and microbial pathogens.
More information: Axel Touchard et al. Deciphering the Molecular Diversity of an Ant Venom Peptidome through a Venomics Approach, Journal of Proteome Research (2018). DOI: 10.1021/acs.jproteome.8b00452
Abstract
The peptide toxins in the venoms of small invertebrates such as stinging ants have rarely been studied due to the limited amount of venom available per individual. We used a venomics strategy to identify the molecular diversity of the venom peptidome for the myrmicine ant Tetramorium bicarinatum. The methodology included (i) peptidomics, in which the venom peptides are sequenced through a de novo mass spectrometry approach or Edman degradation; (ii) transcriptomics, based on RT-PCR-cloning and DNA sequencing; and (iii) the data mining of the RNA-seq in the available transcriptome. Mass spectrometry analysis revealed about 2800 peptides in the venom. However, the de novo sequencing suggested that most of these peptides arose from processing or the artifactual fragmentations of full-length mature peptides. These peptides, called "myrmicitoxins", are produced by a limited number of genes. Thirty-seven peptide precursors were identified and classified into three superfamilies. These precursors are related to pilosulin, secapin or are new ant venom prepro-peptides. The mature myrmicitoxins display sequence homologies with antimicrobial, cytolytic and neurotoxic peptides. The venomics strategy enabled several post-translational modifications in some peptides such as O-glycosylation to be identified. This study provides novel insights into the molecular diversity and evolution of ant venoms.
Journal information: Journal of Proteome Research
Provided by American Chemical Society