Immunotherapy for deadly bacteria shows early promise

If immunotherapy—the harnessing of the body's immune system—can destroy cancer cells, as has been demonstrated, why not try to trigger the body's immune system to battle deadly bacteria?
That question drives Marcos Pires's pursuit of what he calls bacterial immunotherapy or immunobiotics—using the human immune system's powerful mechanisms of preventing entry and colonization of pathogens to defeat the deadliest, antibiotic-resistant bacteria.
Pires and his research team at Lehigh University, where Pires is an Associate Professor in the Department of Chemistry, have previously demonstrated a successful method of labeling the surface of Gram-positive bacteria with antigenic epitopes—the part of a foreign substance that is recognized by the immune system—and then triggering the recruitment of endogenous antibodies.
However, according to Pires, that method was ineffective against Gram-negative bacteria, which have an extra layer of protection around themselves. Gram-negative bacteria—which includes Pseudomonas aeruginosa, associated with serious illnesses such as pneumonia and sepsis, and the food-borne Escherichia coli (E. coli)—are among the hardest bacteria to destroy and the deadliest. These bacteria continually evolve, rendering current antibiotics powerless—and the pipeline for new antibiotic drugs is drying up.
Now, Pires and his team have designed a strategy aimed at tagging Gram-negative bacteria for destruction via small molecule conjugates they have created that specifically home to bacterial cell surfaces and trigger an immune response. The researchers describe their work in a paper to be published in Cell Chemical Biology called: "Synthetic Immunotherapeutics Against Gram-negative Pathogens."
The small-molecule conjugates they have created were assembled using polymyxin B (PMB)—an antibiotic that inherently attaches to the surface of Gram-negative pathogens—and antigenic epitopes that recruit antibodies found in human serum.
"To target these bacteria, we turned to an old class of antibiotics known as colistin," says Pires. "Colistin is a last-resort antibiotic. It just so happens that it destroys bacteria by landing on its surface. We modified colistin with an agent that attracts antibodies onto the surface of the bacteria and built a compound that both directly kills bacteria and at the same time induces an immune-response."
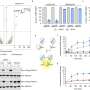
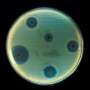
Their compound targets pathogenic bacteria in two distinct ways to generate a very promising lead in immunotherapeutic agents for advanced testing. The team conducted experiments using a panel of Gram-negative pathogens, including E. coli. They treated the bacteria with their compounds in real human serum and observed a significant decrease in the number of live bacteria.
This is a clear indication, says Pires, that the method is working by successfully harnessing the immune system to target this dangerous disease-causing bacteria.
"With this one-two punch against these difficult to kill bacteria, we believe there is great potential for in vivo testing to evaluate them further," says Pires.
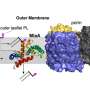
The research brought Pires's group into contact with Lehigh colleague, Wonpil Im, the Presidential Endowed Chair in Health and Professor of Biological Sciences and Bioengineering, in a synergistic, interdisciplinary collaboration. Im, who is a co-author on the paper, uses computational biophysics to learn how antibiotics permeate bacterial membranes and target bacteria for destruction. His research group has developed CHARMM-GUI, an open-access research program that simulates complex biomolecular systems more simply and more precisely than previously possible. The tool is becoming increasingly valuable as more bacteria develop resistance to antibiotics.
"During the optimization stage," says Pires, "we linked up with the Im research group to model how the surface composition of the bacteria may hinder or assist in the immune system activation."
In the paper, the authors write: "In conclusion, we have designed and synthesized a unique class of immunotherapeutic agents that exploits the lipid A binding scaffold of polymyxins to decorate the surface of Gram-negative bacteria with haptens. We showed that the most potent members of this panel trigger the opsonization of E. coli and P. aeruginosa. Most significantly, the lead agent induced CDC-based killing of E. coli. Re- introduction of the membrane-disrupting fatty acid tail restored its inherent antimicrobial activity. Finally, we showed that this agent can target and label the surface of Gram-negative pathogens in a live host. In the future, we plan to expand our in vivo studies to complex animals to establish the suitability of this class of molecules to fight infections. Moreover, we will explore how our strategycan be used to induce the grafting of exogenous haptens onto bacterial cell surfaces with the goal of providing finer control on antibody levels."
More information: Cell Chemical Biology (2018). DOI: 10.1016/j.chembiol.2018.05.019 , www.cell.com/cell-chemical-bio … 2451-9456(18)30190-9
Journal information: Cell Chemical Biology
Provided by Lehigh University