Simulation of the AsqJ enzyme opens up new options for pharmaceutical chemistry
Practically all biochemical processes involve enzymes that accelerate chemical reactions. A research team from the Technical University of Munich (TUM) has now for the first time deciphered the molecular mechanism of the enzyme AsqJ. Their findings might open up new options in the production of pharmaceutically active molecules.
Without enzymes, nature would come to a standstill. These tiny molecules accelerate biochemical reactions or make them possible in the first place. But how does this happen on a molecular level? "Understanding the exact function of enzymes is one of the greatest challenges of modern biochemistry," says Ville Kaila, Professor of Computational Biocatalysis at the Technical University of Munich.
The research team led by Ville Kaila and Michael Groll, Professor of Biochemistry at the Technical University of Munich, have, for the first time, deciphered the mechanism of the enzyme aspoquinolone J (AsqJ), a dioxygenase that activates carbon bonds with oxygen.
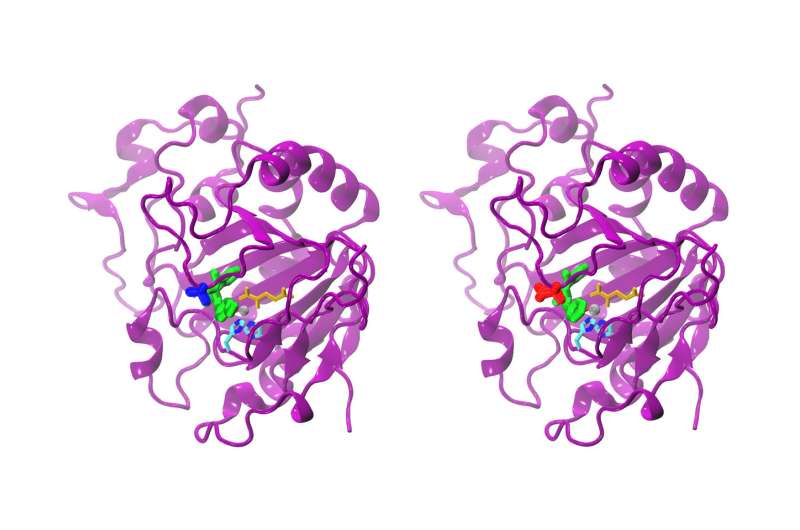
The enzyme AsqJ is particularly exciting as it catalyzes a cascade of chemical reactions that ultimately lead to the formation of antibacterial compounds. It was discovered only a few years ago in the Aspergillus nidulans fungus. The researchers combined different methods to uncover the secrets held within the enzyme: First, Alois Bräuer and Prof. Michael Groll used X-ray crystallography to determine the three-dimensional atomic structure of the molecule. Sophie Mader and Ville Kaila then used this information to carry out quantum mechanical simulations on its biochemical processes.
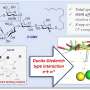
"Our calculations illustrate how the enzyme catalyzes the formation of quinolone alkaloid," reports Kaila. "Tiny details have amazing effects: A slight change in the substrate, like the removal of a small chemical group, is sufficient to practically stop the reaction."
Next, the team computationally designed a new variant of the enzyme that catalyzes the formation of quinolone alkaloids with the modified substrate. This new enzyme was experimentally produced in bacteria and tested for its functionality. "The results were impressive: the expected reaction took place after only a few seconds," recalls Bräuer.
"This experiment demonstrates that our methodology works and is also suited to represent the functionality of other enzymes at the molecular level," says Ville Kaila. Enzyme design is still at a basic level, but it has enormous potential. In the future, we could aim to computationally design medical drugs, for example.
"The work demonstrates that our methodology is accurate and also well suited to study the functionality of other enzymes at the molecular level," says Ville Kaila. Enzyme design is still basic research - but it has enormous potential. An aim of future research will be to design enzymes in a computer to, for example, produce new drugs.
More information: Sophie L. Mader et al, Catalytic mechanism and molecular engineering of quinolone biosynthesis in dioxygenase AsqJ, Nature Communications (2018). DOI: 10.1038/s41467-018-03442-2
Journal information: Nature Communications
Provided by Technical University Munich