Mirror image: Researchers create higher-quality pictures of biospecimens
Researchers from the National Institutes of Health and the University of Chicago improved the speed, resolution, and light efficiency of an optical microscope by switching from a conventional glass coverslip to a reflective, mirrored coverslip and applying new computer algorithms to process the resulting data.
Hari Shroff, Ph.D., chief of the National Institute of Biomedical Imaging and Bioengineering's lab section on High Resolution Optical Imaging (HROI), and his team have spent the last few years developing optical microscopes that produce high resolution images at very high speed. After his lab develops these new microscopes, they release the plans and software for free, so any researcher can replicate the advances made at NIH.
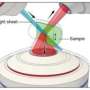
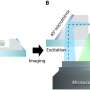
This latest microscope builds on previous improvements that Shroff's lab had made with selective plane illumination microscopy (SPIM). The developments are described in a paper published Nov. 13, 2017, in the advance online edition of Nature Communications. SPIM systems differ from traditional microscopes because they use light sheets to excite the sample, only exposing the imaged sample plane to light. Because only the portion of the sample being imaged (rather than the entire sample) is exposed to light, there is less overall damage to the sample. Thus, SPIM systems are gentler than traditional microscopes.
In 2013, Shroff and his colleague in the HROI lab, Yicong Wu, developed the diSPIM—a SPIM system equipped with two lenses so it obtains two views of the sample instead of just one. Just as using two eyes provides much better depth and three-dimensional perception than using only one eye, the dual view microscope enables 3-D imaging with much greater clarity and resolution than traditional single-view imaging. In 2016, they added a third lens, showing that this additional view can further improve light efficiency and resolution in 3-D imaging.
"Once we incorporated three lenses, we found it became increasingly difficult to add more," said Shroff. "Not because we reached the limit of our computational abilities, but because we ran out of physical space."
The lenses used to image the samples are bulky and need to be close to the samples to clearly image the detailed subcellular structure within a single cell or the neuronal development within a worm embryo. The space around the sample becomes more and more limited with each additional lens.
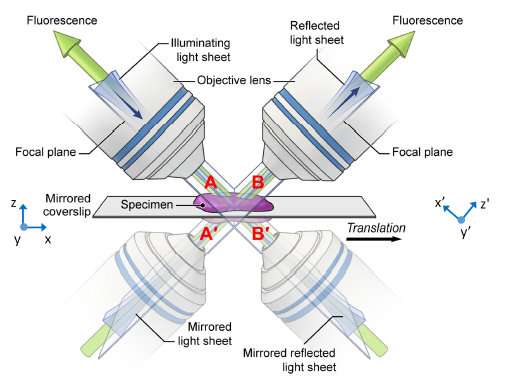
Wu and Shroff's solution was conceptually simple and relatively low-cost. Instead of trying to find ways to stuff in more lenses, they use mirrored coverslips.
"It's a lot like looking into a mirror," Shroff explained. "If you look at a scene in a mirror, you can view perspectives that are otherwise hidden. We used this same principle with the microscope. We can see the sample conventionally using the usual views enabled by the lenses themselves, while at the same time recording the reflected images of the sample provided by the mirror."
One complication is that both the conventional and reflected views contain an unwanted background generated by the light source. In order to deal with this problem, Wu and Shroff collaborated closely with Patrick La Riviere's group at the University of Chicago. La Riviere is an expert in computational imaging, and helped the team create computer processing software that can identify and remove the unwanted background and clarify the image.
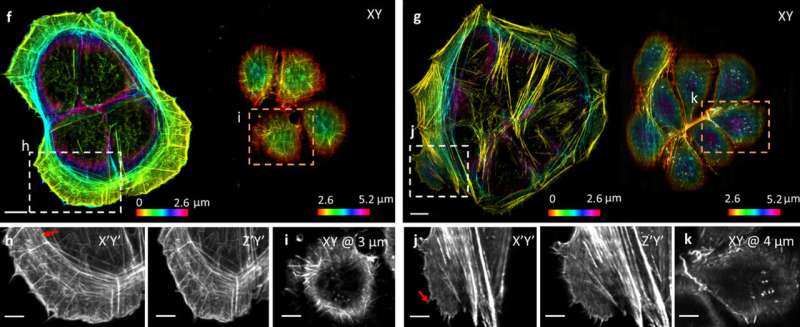
Using the mirrored coverslips in conjunction with the computer software, the team was able to improve the speed two-fold and almost double the resolution in comparison with conventional diSPIM systems without changing the hardware of the microscope. An additional benefit of the technique is that with mirrored coverslips, the microscope is able to collect more light from the sample without increasing the overall light exposure to the sample. As a result, it increases the efficiency by two to three times compared with diSPIM. The researchers hope that in the future this technique may be adapted to other forms of microscopy.
More information: Yicong Wu et al, Reflective imaging improves spatiotemporal resolution and collection efficiency in light sheet microscopy, Nature Communications (2017). DOI: 10.1038/s41467-017-01250-8
Journal information: Nature Communications
Provided by National Institutes of Health