Jellyfish fluorescence shines new light on DNA copying

Scientists at the University of York have used florescent proteins from jellyfish to help shed new light on how DNA replicates.
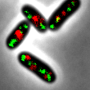
Using these proteins, originally found in jellyfish to make them glow, the team where able to focus laser beams on the brightly lit proteins and track them inside a bacteria that normally lives inside the human gut.
This allowed scientists to watch the molecular machinery of DNA as it replicated inside a cell one molecule at a time. It revealed for the first time that only one component of this process, called DnaB helicase, remains stable - like a molecular anchor to the process.
In most cells, whether human or bacterial, a new cell is created after an existing cell divides in two. This means that a copy of the original sequence of genes coded in its DNA must be precisely copied and placed into the new cell.
This is thought to be a process that occurs slowly and methodically at set points in time. New research at the University of York, in collaboration with the University of Oxford and McGill University Canada, however, has now tracked this replication process in real-time and shown that it is far more dynamic than the textbooks suggest, occurring instead through a 'stuttering-like process' in short bursts.
Pioneering
Professor Mark Leake, Chair of Biological Physics at the University of York, said: "We pioneered a new method of light microscopy which allowed us to see this fascinating replication process occur molecule-by-molecule.
"We were surprised to find, however, that rather than the organised and methodical way that we expected this process to unfold, it instead happened in a 'stuttering' action, much like driving too slowly in high gear of a car. The big question, of course, was why the cell performs an essential process in such an unstable way?
"The stuttering action provide 'checkpoints' at various stages of the DNA copying process to make sure there is no errors made and, if there is, correct them before it is too late. This means that the cells can pause to fix an error in a small fragment of the DNA rather than attempt an unmanageable correction in one complete and huge strand of it.
"Although the process looks inelegant and almost random, it is actually highly efficient."
Human health
The process of DNA replication is fundamental to all life and the way errors in the process are resolved is especially important to human health. Errors can give rise to forms of cancer and become more prevalent in an ageing population.
This work will help scientists not only understand more fully the basic building blocks of life but potentially also provides new insights into a range of health conditions as well as even shedding new light on how human ageing can give rise to diseases associated with errors in copying the DNA from cell to cell.
Research was conducted using the DNA of Escherichia coli cell, bacteria, but However, the next stage of this research will investigate the same process in more complex cells, ultimately including those from humans.
The research, 'Frequent exchange of DNA polymerase during bacterial chromosome replication', was supported by the BBSRC and is published in the journal, eLife.
More information: Thomas R Beattie et al. Frequent exchange of the DNA polymerase during bacterial chromosome replication, eLife (2017). DOI: 10.7554/eLife.21763
Journal information: eLife
Provided by University of York