Computers learn to recognize molecules that can enter cells
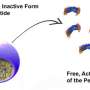
A team of researchers from UCLA and the University of Illinois at Urbana-Champaign originally set out to discover and design antimicrobial peptides—short chains of amino acids that can kill bacteria by punching holes in their cell membranes.
To do this, they developed a computer program that could differentiate between amino acid sequences that can kill bacteria and those that cannot. However, along the way, the researchers also discovered that the program started to recognize features of peptides that could alter the shape of membranes. This shape-altering feature helps peptides travel through the membrane and into the cell, making it possible for the peptides to carry and deliver medicines directly into diseased cells.
Using a screening method, researchers not only found new peptides that had this property, but they also discovered that many known human proteins, longer chains of amino acids, also had this ability, even though membrane-crossing is not their primary known function. This discovery, published in the Proceedings of the National Academy of Sciences, has a wide range of applications in biomedicine, such as combatting infections and delivering drugs directly into cells.
The UCLA researchers, led by Gerard Wong, a professor of bioengineering, spearheaded the experimental work, while the computational tools were developed in collaboration with Andrew Ferguson, a professor of materials science and engineering at Illinois.
"Using machine learning, we developed a computer program that can differentiate between a peptide sequence that is antimicrobial and one that isn't antimicrobial," said Wong, the co-senior author of the project and a member of the California NanoSystems Institute at UCLA. "During this process, we serendipitously discovered a way to differentiate between peptides that permeate membranes and peptides that don't."
They studied a class of well-known peptides called antimicrobial peptides, which are proteins that help the immune system by killing bacteria primarily by permeating the membrane.
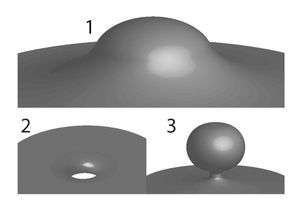
During their experiments, the researchers found that the program, originally created to recognize anti-microbial peptides, was simultaneously finding peptides that generate saddle-shaped curvature on the cell membrane. This curvature is much more conducive to allowing a puncture through the membrane into the cell.
Using this new tool, the researchers performed a search of possible peptide sequences to find new membrane-active peptides that do not occur naturally, but can be created chemically.
"Not only are we able to design better antimicrobials to combat drug-resistant infections, we can also design peptides to deliver medicines or other cargo into cells, understand how viruses and bacteria bypass membranes, and uncover membrane activity in proteins that previously have not been characterized to have membrane activity," said Ernest Lee, the paper's lead author and a student in the UCLA-Caltech Medical Scientist Training Program.
More information: Ernest Y. Lee et al. Mapping membrane activity in undiscovered peptide sequence space using machine learning, Proceedings of the National Academy of Sciences (2016). DOI: 10.1073/pnas.1609893113
Journal information: Proceedings of the National Academy of Sciences
Provided by University of California, Los Angeles