Rock meets microbe: Towards better models of subsurface microbial communities
Beneath the land are subsurface aquifers in which deposits of gravels, sands, silts, and clays mix with water and microbial communities. These subsurface gatherings of microbes, metabolically influenced by the mineral mixes that bind them, are busy with the business of microscopic life: eating carbon and excreting gases, which in turn (by virtue of microbial abundance) greatly influence the composition of the Earth's atmosphere.
Data about microbial communities, even those underwater or in the transition zones between water and land, are important. They inform models designed to predict climate and other large-scale phenomena that are consequential for the planet.
These subsurface microbial communities, especially in key transition zones between water and land, are hard to access and hard to study, so modelers need proxy variables to predict their likely spatial distribution. In a recent paper, a team of researchers led by James C. Stegen and Jim K. Fredrickson at PNNL describe an unexploited opportunity for modeling the distribution of subsurface microbial communities. If the mineralogy and other characteristics of sediments influence subsurface microbial communities, the researchers reasoned, then spatial distributions of sediment characteristics - which are easier to describe - can be used to predict the spatial distribution of biogeochemically relevant microbial communities.
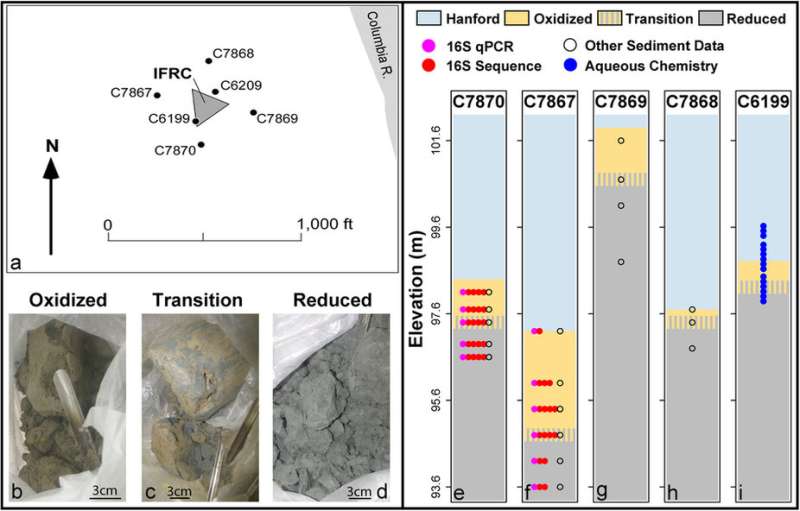
It's like getting rocks to talk, or at least show pictures. (Figure 8 in the paper is a three-dimensional display showing how microbial biomass likely varies through the subsurface due to spatial variation in sediment properties.) Along a stretch of the Columbia River near Richland, Wash., Stegen and his team worked at the Hanford Site 300 Area, where the subsurface geochemical and biogeochemical processes that influence the transport of contaminants have already been widely identified. They also took advantage of 35 extant boreholes and the strongly vertical (and weakly horizontal) structure of the area's fine-grained Ringold geologic formation.
The researchers focused on three biogeochemical conditions within the Ringold formation, which they characterized as "oxidized," "reduced," or "transition" biogeochemical facies. The facies concept provides a way to categorize sediment properties into discrete bins. While it is often used to describe "lithofacies" based on sediment physical properties, here the authors applied this approach to biogeochemical properties.
Previous papers have used hydrogeological properties as proxies for microbial activity. But none (until now) have leveraged biogeochemical facies in order to spatially project microbial biomass. Such an approach, the authors said, will generate fundamental knowledge about the spatial distributions of key properties within microbial communities. It will also provide important constraints to hydro-biogeochemical models in both initial and dynamic conditions.
The researchers set out to evaluate their hypotheses: that the richness of a microbial community will most strongly be related to redox state, which influences how much energy is available to microbial cells. And that mixing complementary electron donors and acceptors will create a biogeochemical "hot spot" in the transition zone, resulting in elevated microbial biomass. From there, the researchers coupled variations in microbial biomass with spatial distributions of biogeochemical facies to predict microbial biomass across a 3-dimensional spatial domain.
Along the way, they ran into some surprises. An abundance of organic carbon, for instance, did not equate to higher numbers of microbial species. That deviates from macro-ecological observations that equate increases in taxonomic richness with increases in energy supply.
In the field, the team sonically drilled four hydrological monitoring wells and recovered sediments from multiple depths. They carefully transported their samples to a laboratory setting on wet ice and in anaerobic glove bags in order to maintain stable redox conditions.
From there, researchers employed multiple methods to characterize the sediment samples. Some were dried and ground to be analyzed by X-ray diffraction; others were impregnated with epoxy and imaged using X-ray tomography. Other samples were assessed for sulfide mineralization. Additional material was used to extract DNA, which subsequently helped characterize both microbial community composition (via sequencing) and microbial biomass. Groundwater was sampled and analyzed across a vertically structured redox transition zone that aligned with the sediment samples.
To arrive at a 3-dimensional map of microbial biomass, the researchers used geologist well logs from each of the 35 boreholes in order to define elevations of the redox transition zones, where they expected to find the highest microbial biomass. They used these data to generate a 3-D reconstruction of biogeochemical facies, combining that with facies-specific microbial biomass estimates. In turn, the researchers were able to generate a 3-D map of microbial biomass, based on the more easily measured proxy variable of redox state.
They say future hydro-biogeochemical simulations could be more realistic by adding biomass values to grid cells within each biogeochemical facies. Or by applying this same approach across lithofacies to enable broader predictions of microbial community properties. Although PNNL researchers called for this approach of using proxy variables decades ago, it has received limited attention. The researchers say it has great potential for creating multi-scale models that have field-scale predictive value for biogeochemical function in the Earth's climate-critical transition zones between water and land.
More information: James C. Stegen et al. Coupling among Microbial Communities, Biogeochemistry, and Mineralogy across Biogeochemical Facies, Scientific Reports (2016). DOI: 10.1038/srep30553
Journal information: Scientific Reports
Provided by Pacific Northwest National Laboratory