Novel study on histones provides better understanding of gene regulation
Scientists from A*STAR's Genome Institute of Singapore (GIS) have discovered the unique functions of a specific type of histone modification, which may prove crucial in understanding gene regulation and the development of diseases. This development was published in the scientific journal Genome Research.
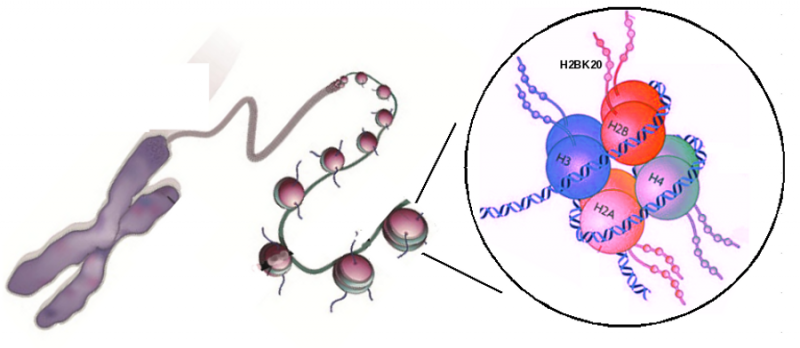
Histones are proteins found in the cell nucleus, which organise DNA into structural units called nucleosomes, helping to protect DNA as well as control gene expression. There are five main families of histones, and they have different kinds of modifications. Researchers observe patterns of these histone modifications across the whole genome in order to understand the process of gene regulation and of disease progression. For example, a kind of histone modification called histone acetylation has been linked to activation of different sites on DNA for gene replication and enhancing transcriptions. However, much remains to be known about histones and how they function. While there are 35 known histone acetylations, many of them are not characterised properly. The majority of regulatory genomics studies also focus exclusively on two histone acetylations known as H3K27ac and H3K9ac.
GIS scientists, led by Dr Shyam Prabhakar, Associate Director of Integrative Genomics, performed an unbiased comparison of different histone modifications, to test their capacity to highlight a set of active enhancers. They identified a histone acetylation known as H2BK20ac as the most predictive marker of active enhancers. Currently there is no extensive study on H2BK20ac properties. The scientists also discovered that among all histone acetylations, H2BK20ac is the most efficient for identifying cell-state specific active promoters, and that this histone modification is absent at promoters of house-keeping genes which are active in all cell-types. They also found that external signalling to cells can cause heterogeneity of occurrences of histone acetylations especially H3K27ac and H2BK20ac.
First author Dr Vibhor Kumar, Research Scientist, Computational and Systems Biology at the GIS said, "Currently, clinicians try to find genes or markers which can provide information on the state of cell, such as whether it is diseased or normal for example. Our finding will help them to find promoters specific to a disease or cell-state more easily because they do not have to perform comparisons with other cell-types."
Prof Keji Zhao, Senior Investigator at the Laboratory of Epigenome Biology, National Institutes of Health, USA, said, "I think it is a significant progress in the field and the results are very important for accurately annotating functional regulatory elements in the genome."
"This well-designed and rigorously conducted study has uncovered new chromatin features of enhancers, an abundant class of regulatory DNA. The finding could help better understand the mechanisms of enhancers, which are still pretty rudimentary today," said Prof Bing Ren, Professor of Cellular and Molecular Medicine, Institute of Genomic Medicine, University of California, San Diego, USA.
GIS Executive Director Prof Ng Huck Hui said, "This is indeed an exciting study because our discovery of the heterogeneity of histone acetylation at active promoters and enhancers has the potential to influence and significantly enhance future epigenome base studies which can advance the development of targeted therapeutic solutions."
More information: Vibhor Kumar et al. Comprehensive benchmarking reveals H2BK20 acetylation as a distinctive signature of cell-state-specific enhancers and promoters, Genome Research (2016). DOI: 10.1101/gr.201038.115
Journal information: Genome Research