May 10, 2016 weblog
Procedure developed to decode histone changes that point out gene regulation
(Phys.org)—A team of researchers with Massachusetts General Hospital, Harvard Medical School and the Broad Institute has developed a procedure to decode histone modifications that point out gene regulation. In their paper published in the journal Science, the team describes the approach they developed to map combinations of modifications made to individual nucleosomes.
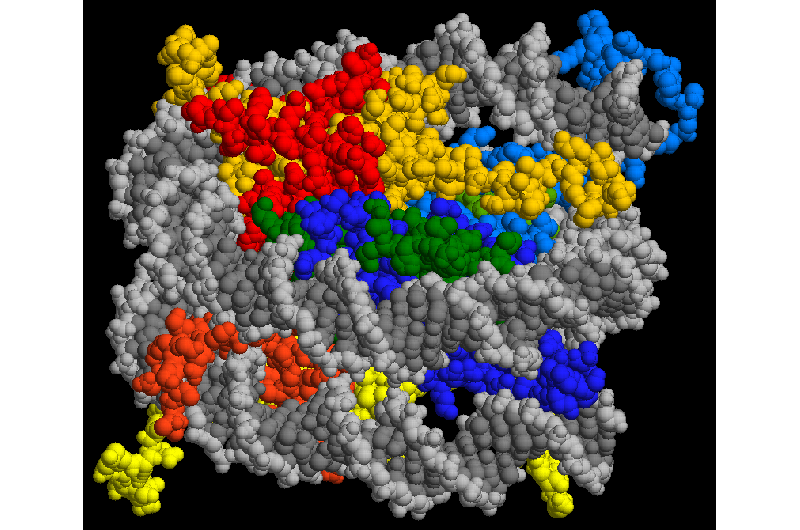
Determining whether a gene should be active or not has been linked to covalent modification of histone proteins that form the framework upon which DNA strands are wound. Scientists would like to better understand how the coding works but have not been able to find a way to uncover the combinations at play, or to quantify or map them. In this new effort, the researchers report that they have developed a technique that allows for doing just that. Their strategy involved starting with pluripotent stem cells as well as lineage-committed cells, singling out nucleosomes (the packages that contain histone proteins and DNA strands) and dabbing the ends of DNA strands with fluorescent material, meanwhile the DNA was incubated with histones that had been flagged with fluorescent antibodies.
Next to pinpoint individual nucleosomes, the team used internal reflection microscopy which allowed for creating millions of images which set the stage for decoding modification states. In so doing, the team was able to actually quantify different changes on single nucleosomes, which led to demonstrating that genetic and chemical changes preferentially impacted nucleosomes with certain modification states. The last step involved applying the same approach to single-molecule DNA sequencing to figure out the exact location of the modified nucleosomes in a given genome.
In using their technique, the team reports that they found evidence that suggested that cells that had been differentiated showed different patterns of "bivalent" markings than did embryonic cells. They suggest that the technique holds promise of assisting in answering some of the fundamental questions in both chromatin biology and epigenetic regulation by providing new insights into genetic processes. They also note that their results represent early results in their efforts to better understand histone signaling regulation and expect that eventually they will have a more complete understanding of the rules that underlie the fate of cells in general.
More information: E. Shema et al. Single-molecule decoding of combinatorially modified nucleosomes, Science (2016). DOI: 10.1126/science.aad7701
Abstract
Different combinations of histone modifications have been proposed to signal distinct gene regulatory functions, but this area is poorly addressed by existing technologies. We applied high-throughput single-molecule imaging to decode combinatorial modifications on millions of individual nucleosomes from pluripotent stem cells and lineage-committed cells. We identified definitively bivalent nucleosomes with concomitant repressive and activating marks, as well as other combinatorial modification states whose prevalence varies with developmental potency. We showed that genetic and chemical perturbations of chromatin enzymes preferentially affect nucleosomes harboring specific modification states. Last, we combined this proteomic platform with single-molecule DNA sequencing technology to simultaneously determine the modification states and genomic positions of individual nucleosomes. This single-molecule technology has the potential to address fundamental questions in chromatin biology and epigenetic regulation.
Journal information: Science
© 2016 Phys.org