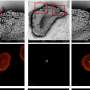
AFM films 'living' nuclear pore complexes at work for the first time

Using an ultra fast-scanning atomic force microscope, a team of researchers from the University of Basel has filmed "living" nuclear pore complexes at work for the first time. Nuclear pores are molecular machines that control the traffic entering or exiting the cell nucleus. In their article published in Nature Nanotechnology, the researchers explain how the passage of unwanted molecules is prevented by rapidly moving molecular "tentacles" inside the pore.
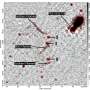
The atomic force microscope (AFM) is not a microscope to look through. Like a blind man uses his fingers, it "feels" a surface with an extremely fine tip to resolve tiny cellular structures of only millionths of a millimeter in size, such as the pores in the nuclear envelope. However, this process is normally slow and can take up to one minute to capture an image. In comparison, modern high-speed AFMs are able to record movies of molecules in action by capturing several hundred images per minute.
Using high-speed AFM, Roderick Lim, Argovia Professor at the Biozentrum and the Swiss Nanoscience Institute of the University of Basel, has not only directly visualized the nuclear pore's selective barrier, but also its dynamic behavior to resolve a long-standing mystery of how unwanted molecules are prevented from entering the nucleus.
Nuclear pore complexes regulate the transport of molecules
The overall structure of the nuclear pores is generally known. These are not simple holes, but are massive transport hubs that incorporate by the thousands into the nuclear membrane. They have a donut-shaped structure consisting of about thirty different proteins, called nucleoporins, and a central transport channel. Within the pore, several disordered proteins (FG Nups) form a selectivity barrier or filter. While small molecules can easily pass this barrier, large molecules such as proteins are prevented from entering the nuclear pore. An exception to this are the proteins needed in the cell nucleus, for example, for the repair or replication of genetic material. Their translocation from the cytoplasm to the nucleus is assisted by transport receptors that recognize a specific "address tag" carried by these proteins.
High-speed AFM reveals dynamic processes
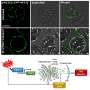
"With the high-speed AFM we could for the first time, peer inside native nuclear pore complexes, only forty nanometers in size", says Lim. "This method is a real game changer. We could see the individual FG Nups and film them in action. This wasn't possible until now!"
Additionally, Yusuke Sakiyama, the PhD student who performed the experiments, had to grow super-sharp carbon nanofibers on each high-speed probe in order to reach inside the NPC. This then generates a video sequence from multiple images that enables the researcher to observe the "true to life" dynamics of biological processes at the nanometer level.
A barrier of undulating molecular "tentacles"
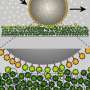
Due to the high spatial and temporal resolution, the scientists were able to show that the FG Nup filaments are highly flexible. "They are not stiff bristles but quite the contrary. Like the thinnest tentacles, the FG Nups rapidly fluctuate, elongate and retract, and sometimes even briefly intermingle within the pore", says Lim. The speed of their motion determines which molecules can pass through the pore. "Large particles move much more slowly than the FG Nups and are thus hindered from entering the NPC by repeated collisions", explains Lim. "Small molecules, however, undergo rapid diffusion and have a high probability of passing the FG Nup barrier."
By understanding how NPCs function as transport hubs in living cells, Lim who is a member of the NCCR Molecular Systems Engineering is now investigating how NPC-inspired selective filters might regulate molecular traffic in non-biological systems.
More information: Yusuke Sakiyama et al, Spatiotemporal dynamics of the nuclear pore complex transport barrier resolved by high-speed atomic force microscopy, Nature Nanotechnology (2016). DOI: 10.1038/nnano.2016.62
Journal information: Nature Nanotechnology
Provided by University of Basel