Simple mechanism may have allowed primitive cells to maintain internal conditions
A Massachusetts General Hospital (MGH) research team investigating how the earliest stages of life might have developed has discovered a way the first living cells could have met a key challenge—maintaining a constant internal environment, a process called homeostasis, even when external conditions change. In a report receiving advance online publication in Nature Chemistry, the investigators describe finding that the dilution of a simple cell's internal contents that takes place when the cell grows increases the activity of an internal enzyme, maintaining activity at a constant level relative to cellular volume.
"Modern cells are constantly regulating what they are doing—synthesizing, degrading and exporting a whole suite of RNAs and proteins—depending on the cell's particular needs at the time," says Aaron Engelhart, PhD, of the MGH Department of Molecular Biology and the Center for Computational and Integrative Biology, lead author of the paper. "One would expect that the earliest cells weren't nearly as complex as today's cells, but they still had the need to regulate their internal environment. The sort of regulation we've shown here—switching on enzymes during growth—is perhaps the simplest form of the internal regulation that a primitive cell might have needed."
Engelhart is a postdoctoral fellow in the laboratory of Jack Szostak, PhD, senior author of the report. A co-recipient of the 2009 Nobel Prize in Physiology or Medicine for his contribution to the discovery of the enzyme telomerase, Szostak and his team have been investigating the origins of life for more than 15 years, and much of their work has focused on developing a model protocell—a primitive, synthetic cell consisting of nucleic acid strands enclosed by a membrane that would be capable of growth, replication and evolution.
Previous studies have shown that simple membranes comprised of fatty acids and lacking the complex molecular components of modern cells would still be permeable to small nutrient molecules, including those needed to assemble nucleic acids, such as RNA or DNA. A 2013 study showed it was possible to copy molecules of RNA—which many believe was the genetic blueprint of the first cells—without the complex enzymes used by today's cells, even within small sacs or vesicles formed from fatty acid membranes. Szostak's team has also found that such vesicles will expand under certain conditions, raising the question of how vesicles' internal environment can be maintained, since their contents would become diluted as the enclosing membranes expand.
To investigate this question, Engelhart and his colleagues examined how the concentration of a protocell's contents might affect the activity of RNA enzymes, called ribozymes. They hypothesized that high levels of small nucleic acid strands within a cell might bind to corresponding sequences on a ribozyme, suppressing its activity. They first tested this in free solution—not within vesicles—and found that short RNA strands could totally shut down ribozyme activity at high concentrations but had little effect at low concentrations.
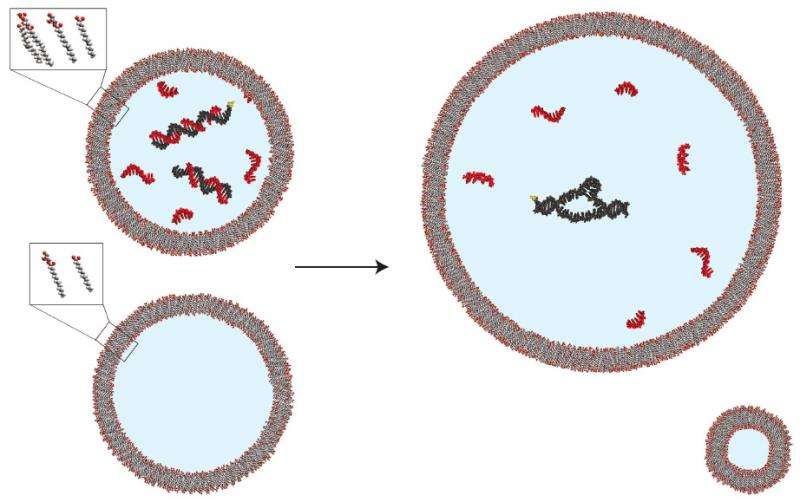
Experiments with vesicles containing both ribozymes and short strands of RNA specifically designed to bind to the ribozymes showed that enzymatic activity remained steady as the vesicles expanded; but in vesicles containing the enzymes alone, activity dropped as the vesicles grew. The researchers also observed this regulatory effect when the short strands of RNA contained random sequences.
"Without some sort of regulatory mechanism like we've shown here, cellular growth would be accompanied by some loss of function, since active enzymes would be present at a lower concentration in larger cells," says Engelhart. "We haven't extended this work to dividing vesicles yet, but a key long-term goal of the lab is developing a full, chemically based primitive cell cycle, including both growth and replication. A number of people in the lab are working on the next step towards that—systems that will allow us to make many copies of a single strand of RNA—and it will be very exciting to see how systems like the one we've explored in this paper work in such a cycle."
More information: A simple physical mechanism enables homeostasis in primitive cells, Nature Chemistry, DOI: 10.1038/nchem.2475
Journal information: Nature Chemistry
Provided by Massachusetts General Hospital