Sophisticated computer simulation shows image of molecular dynamics in cells

While many people may think of cholesterol only as the culprit in heart disease and other serious conditions, this miraculous molecule is actually essential to our cells—organizing the fatty acids in the membranes that surround cells to make those barriers stronger.
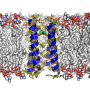
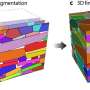
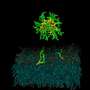
The specifics of how that organizing happens are not well understood, but a research team including a University of Delaware physicist has published a study that shows in colorful detail a computer simulation of a mixture of two lipids and cholesterol—a simple model of a cell membrane.
The configuration of the mixture, sampled by a computer simulation, is the illustration on the cover of the Sept. 1 Biophysical Journal.
The journal article is co-authored by Edward Lyman, assistant professor of physics and astronomy and of chemistry and biochemistry at UD, and Alexander J. Sodt and Richard W. Pastor of the National Heart, Lung and Blood Institute of the National Institutes of Health (NIH).
The cover image illustrates the use of molecular dynamics, or computer simulations of the movements of atoms and molecules. Carrying out the kind of simulation discussed in the Biophysical Journal paper was not an easy task, Lyman said, adding that it would have been too time-consuming for even a standard supercomputer.
"Molecular dynamics is basically looking at balls and springs in a box," Lyman said, visualizing a molecule as composed of atoms (balls) connected by bonds (springs). "And in this case, we've got a lot of balls and springs, so the simulation requires a specialized kind of computer designed explicitly for this type of calculation.
"To do the calculations we did for this paper, you'd have to run a normal supercomputer for about a year. With access to the specialized computer, it took us about a day."
The special-purpose supercomputer the researchers used, called "Anton," was designed and built for molecular dynamics calculations by D.E. Shaw Research and was donated to the scientific community for academic research. Anton is housed and maintained by the Pittsburgh Supercomputing Center, with a staff supported by the NIH.
Lyman's team developed a series of successful research proposals and was awarded time on Anton in three consecutive years.
Although the simulations involved molecular dynamics, and not such medical concerns as how diet or medications can lower a person's blood cholesterol levels, Lyman said the issues are related.
"Cholesterol has many roles in the membranes of your cells, one of which is to organize the cellular machinery responsible for cell-to-cell communication," he said. "Changes in membrane organization change the way cells talk to each other, which is of fundamental importance to understanding both healthy and disease states. So, while we are not studying heart disease directly, we are interested in how gross changes to your cell membranes translate into health outcomes."
Further research, conducting simulations of even more complex mixtures, might examine how membrane organization changes when a person's diet changes. The eventual goal is to leverage the research results to develop therapeutic treatments, Lyman said, in close partnership with experimental collaborators.
Such work is already underway in collaboration with groups at Tulane University, the University of Southern California and the University of Texas Medical School.
"Whatever is revealed, it is clear that cholesterol will play a leading role," the researchers wrote on the official blog of the Biophysical Society.
More information: "Hexagonal Substructure and Hydrogen Bonding in Liquid-Ordered Phases Containing Palmitoyl Sphingomyelin." DOI: dx.doi.org/10.1016/j.bpj.2015.07.036
Journal information: Biophysical Journal
Provided by University of Delaware