Researchers bring to life proteins' motion
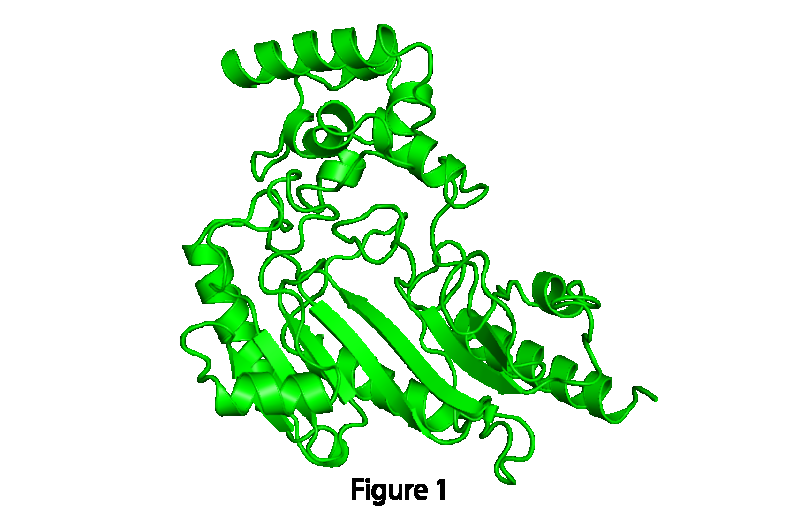
Advancing the field of structural biology that underpins how things work in a cell, researchers have identified how proteins change their shape when performing specific functions. The study's fresh insights, published online in the journal Structure, provide a more complete picture of how proteins move, laying a foundation of understanding that will help determine the molecular causes of human disease and the development of more potent drug treatments.
Though it has long been recognized that proteins are not static, for more than 30 years, scientists' understanding of proteins was limited to observing their structure through static snapshots taken using crystallography, a technique used to capture images of molecules' structures in various configurations.
In order to understand proteins' normal functioning, and design treatments for when cells malfunction, scientists needed, but lacked, a dynamic picture of proteins' coordinated motions. The one experimental technique through which scientists could observe motion, nuclear magnetic resonance (NMR) spectroscopy, has been used with fewer than 1 percent of proteins because the majority of proteins are too large, complex, difficult to obtain or not soluble.
To fill the gap in knowledge, the researchers designed a computer method that looked at two different snapshots of the same protein structures, captured through crystallography, and identified hinge points, or 'hot spots," of flexibility. In doing so, the researchers successfully affirmed the premise that proteins use the ranges of motion that are most efficient. Through improved understanding of the mechanics of motion, researchers will be better able to analyze the changes in proteins as they interact with other molecules, regulating how cells function.
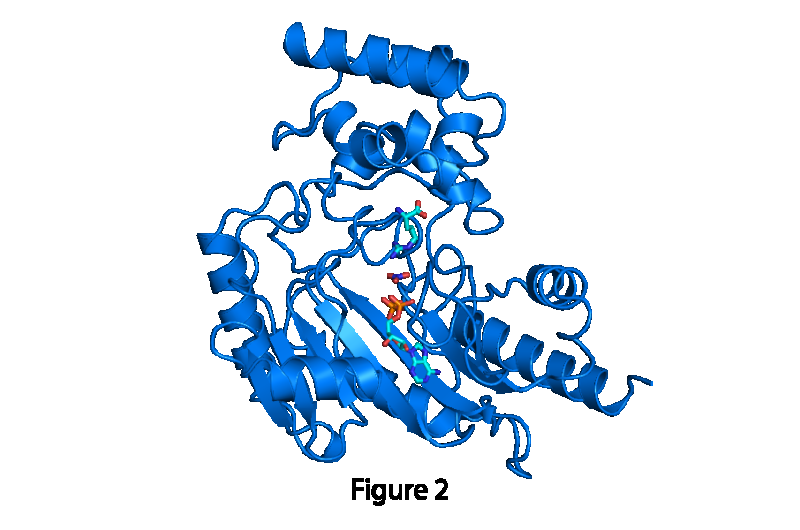
"For more than 30 years, the biology textbooks of medical students have contained static pictures of the structures of more than 100,000 proteins. This study is a step toward bringing those snapshots to life by understanding the motions that are involved in how proteins catalyze reactions or regulate biological processes," said Michael Chapman, Ph.D., professor of biochemistry and molecular biology, OHSU School of Medicine, and member of the Knight Cancer Institute. "By way of analogy, a snapshot of a thoroughbred horse doesn't tell you much about how it's running. This paper is a step toward learning how proteins are moving to go about their business."
More information: Parsimony in Protein Conformational Change, Structure, 2015.
Journal information: Structure
Provided by Oregon Health & Science University