Odd histone helps suppress jumping genes in stem cells, study says

A family of proteins known as histones provides support and structure to DNA, but for years, scientists have been puzzling over occasional outliers among these histones, which appear to exist for specific, but often mysterious reasons. Now, researchers have uncovered a new purpose for one such histone variant: preventing genetic mutations by keeping certain so-called "jumping genes" in place.
This research, which began at Rockefeller University and was published May 4 in Nature, reveals a basic mechanism by which epigenetics, or the control of inherited traits through means other than DNA, works. Due to histones' close relationship with DNA, scientists have known for some time that they are frequently involved in epigenetic control of genes. In this case, one particular histone variant appears to reduce the chance of potentially harmful changes in the stem cells that will eventually generate the various types of tissue that make up a living creature.
"They say that good things come in small packages. Nowhere is this more true than with histone variants. This study found the variant H3.3, which differs only slightly from the standard H3 histones, helps prevent certain genetic elements, which are remnants left behind by ancient viral infections, from moving about within the genome," says study author C. David Allis, Joy and Jack Fishman Professor and head of the Laboratory of Chromatin Biology and Epigenetics. "This discovery is an important addition to our still-evolving knowledge of how epigenetics works at the molecular level."
Histones are proteins that act as spools for the thread that is DNA, giving it support and structure. Chemical modifications to these histones can change the expression of genes, making them more available for expression or silencing them by compacting the DNA-protein complex. Oddball H3.3 varies from its regular counterpart H3 by only few amino acids. Because it is present throughout the animal kingdom, however, scientists have suspected for some time that H3.3 has a specific biological role.
Study authors Simon Elsasser and Laura Banaszynski, both of whom worked on H3.3 in Allis's lab at Rockefeller but have since moved on to other institutions, started by looking at the locations on the mouse genome where H3.3 was deposited in stem cells. Elsasser began the project as graduate student in Allis's lab and continued as a postdoc at the MRC Laboratory of Molecular Biology in the United Kingdom. He is now an assistant professor at the Karolinska Institute in Sweden. He had the idea to look for H3.3 at repetitive sequences; however, repeats are normally filtered out in a genome-wide study. So, Elsasser developed a new approach to capture this information.
A pattern emerged from the results: H3.3 appeared at a certain type of repetitive sequence: retrotransposons, which are leftovers from ancient viral infections. Unlike their ancestral viruses, retrotransposons are trapped in the host genome, but they can still copy themselves and jump to new locations within it. Sometimes, evolution finds a use for them. For instance, retrotransposon-derived genes code for proteins necessary for the placenta in mammals. But when retrotransposons jump, they can also cause harmful mutations.
For studies like this one, which explores chromatin's role regulating gene expression, scientists often use mouse embryonic stem cells. Stem cells' chromatin landscape is more plastic than that of differentiated cells, reflecting their capacity to enter any of many gene expression programs that lead to the hundreds of different cell types in an adult organism. Once the cells have begun to pick an identity, parts of the genome not needed for that identity get closed off forever. Prior to the current study, scientists knew mouse stem cells kept most of the genome accessible, while keeping the lid on retrotransposons by tagging them with chemical markers containing three methyl groups on histone H3.
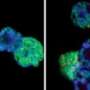
Early experiments done by Banaszynski, while a postdoc in Allis's lab, suggested that H3.3 is necessary for the placement of these suppressive "trimethyl" marks. "By taking away proteins responsible for placing H3.3 into chromatin, or eliminating H3.3 completely, we confirmed that trimethylation depends on H3.3," says Banaszynski, who is currently an assistant professor at the University of Texas Southwestern Medical Center.
"Furthermore, retrotransposons became more active in cells without H3.3, and in these cells, we saw chromosomal abnormalities. It may be that by silencing retrotransposons, H3.3 prevents these abnormalities, however we cannot eliminate the possibility that loss of H3.3 results in this genomic instability for other reasons," Elsasser says.
Although the types of retrotransposons studied in these experiments are not active in humans, it's likely that human stem cells do use H3.3 to keep other varieties of jumping genes in place, Banaszynski says.
The research has implications beyond epigenetics. "This study also hints at a fascinating question in biology: How do cells balance the potential evolutionary benefit of mobile elements, such as retrotransposons, with the competing need to silence them so as to maintain the genome?" she says.
More information: Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells, Nature, DOI: 10.1038/nature14345
Journal information: Nature
Provided by Rockefeller University