Researchers engineer E. coli to produce new forms of popular antibiotic
Like a dairy farmer tending to a herd of cows to produce milk, researchers are tending to colonies of the bacteria Escherichia coli (E. coli) to produce new forms of antibiotics—including three that show promise in fighting drug-resistant bacteria.
The research, which will be published May 29 in the journal Science Advances, was led by Blaine A. Pfeifer, an associate professor of chemical and biological engineering in the University at Buffalo School of Engineering and Applied Sciences. His team included first author Guojian Zhang, Yi Li and Lei Fang, all in the Department of Chemical and Biological Engineering.
For more than a decade, Pfeifer has been studying how to engineer E. coli to generate new varieties of erythromycin, a popular antibiotic. In the new study, he and colleagues report that they have done this successfully, harnessing E. coli to synthesize dozens of new forms of the drug that have a slightly different structure from existing versions.
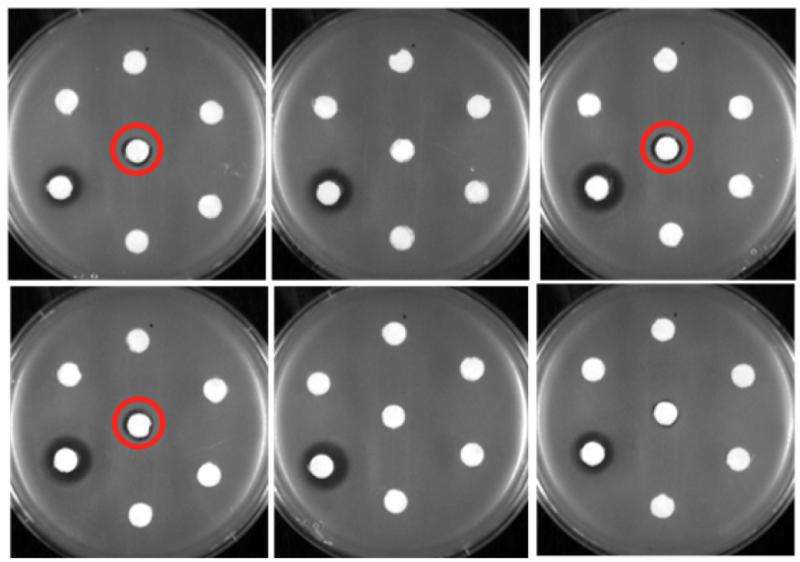
Three of these new varieties of erythromycin successfully killed bacteria of the species Bacillus subtilis that were resistant to the original form of erythromycin used clinically.
'We're focused on trying to come up with new antibiotics that can overcome antibiotic resistance, and we see this as an important step forward,' said Pfeifer, Ph.D.
'We have not only created new analogs of erythromycin, but also developed a platform for using E. coli to produce the drug,' he said. 'This opens the door for additional engineering possibilities in the future; it could lead to even more new forms of the drug.'
The study is especially important with antibiotic resistance on the rise. Erythromycin is used to treat a variety of illnesses, from pneumonia and whooping cough to skin and urinary tract infections.
E. coli as a factory
Getting E. coli to produce new antibiotics has been something of a holy grail for researchers in the field.
That's because E. coli grows rapidly, which speeds experimental steps and aids efforts to develop and scale up production of drugs. The species also accepts new genes relatively easily, making it a prime candidate for engineering.
While news reports often focus on the dangers of E. coli, most types of this bacteria are actually harmless, including those used by Pfeifer's team in the lab.
Over the past 11 years, Pfeifer's research has focused on manipulating E. coli so that the organism produces all of the materials necessary for creating erythromycin. You can think of this like stocking a factory with all the necessary parts and equipment for building a car or a plane.
With that phase of the research complete, Pfeifer has turned to the next goal: Tweaking the way his engineered E. coli produce erythromycin so that the drug they make is slightly different than versions used in hospitals today.
That's the topic of the new Science Advances paper.
The process of creating erythromycin begins with three basic building blocks called metabolic precursors—chemical compounds that are combined and manipulated through an assembly line-like process to form the final product, erythromycin.
To build new varieties of erythromycin with a slightly different shape, scientists can theoretically target any part of this assembly line, using various techniques to affix parts with structures that deviate slightly from the originals. (On an assembly line for cars, this would be akin to screwing on a door handle with a slightly different shape.)
In the new study, Pfeifer's team focused on a step in the building process that had previously received little attention from researchers, a step near the end.
The researchers focused on using enzymes to attach 16 different shapes of sugar molecules to a molecule called 6-deoxyerythronolide B. Every one of these sugar molecules was successfully adhered, leading, at the end of the assembly line, to more than 40 new analogs of erythromycin—three of which showed an ability to fight erythromycin-resistant bacteria in lab experiments.
'The system we've created is surprisingly flexible, and that's one of the great things about it,' Pfeifer said. 'We have established a platform for using E. coli to produce erythromycin, and now that we've got it, we can start altering it in new ways.'
More information: Tailoring pathway modularity in the biosynthesis of erythromycin analogs heterologously engineered in E. coli, Science Advances, advances.sciencemag.org/content/1/4/e1500077
Journal information: Science Advances
Provided by University at Buffalo