Certain virus-derived genomic elements have now been shown to actively regulate important developmental processes
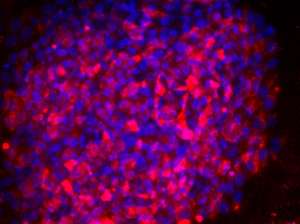
Like fossils buried beneath a modern landscape, the human genome is littered with sequences that originated from ancient viral DNA insertion events. Scientists have long assumed that these 'transposable elements' are, like fossils, biologically inactive and primarily interesting as a window into evolutionary history. However, researchers at the A*STAR Genome Institute of Singapore have now uncovered evidence that some of these sequences play a prominent role in early embryonic development.
Huck-Hui Ng and colleagues embarked on this project in collaboration with Guillaume Bourque at Canada's McGill University. Bourque's group had discovered that one particular class of sequences of transposable elements—known as human endogenous retrovirus subfamily H (HERV-H)—appears to be specifically expressed in human embryonic stem cells (hESCs). Indeed, these HERV-H sequences are actively transcribed in hESCs, producing enigmatic RNA strands that do not encode a protein but nevertheless appear to serve some function.
Ng and Bourque set out to clarify the role of this RNA by performing experiments in which they selectively depleted it from stem cells. hESCs are actively maintained in a so-called 'pluripotent' state, from which they are capable of developing into any cell type in the human body (see image). In the absence of HERV-H RNA, hESCs rapidly lost their pluripotency; the researchers noted that the loss of HERV-H expression considerably altered the activity of many genes associated with cell development and proliferation.
Closer analysis revealed that the HERV-H RNA binds to multiple protein partners, including a protein called OCT4, which plays a role in establishing and maintaining stem cell pluripotency. Ng and Bourque's team determined that the resulting RNA-protein complexes may assemble at specific sites within the HERV-H genomic element. These sequences in turn act as 'enhancers', which help coordinate the activation of other pluripotency-related genes residing in both adjacent and remote regions of the genome.
According to the study's lead author, Xinyi Lu, a postdoctoral researcher in Ng's laboratory, the emergence of the regulatory activities executed by HERV-H could represent an important step in the evolution of our early ancestors. "HERV-H first integrated into the primate genome around 45 million years ago and is only found in the primate genome," says Lu, "and so it may contribute to some of the differences between primates and other mammals."
In future work, Ng and colleagues hope to extend their investigation of HERV-H function to identify other possible protein partners and perhaps other mechanisms by which these ancient sequences exert their influence.
More information: Lu, X., Sachs, F., Ramsay, L., Jacques, P.-É., Göke, J., Bourque, G. & Ng, H.-H. "The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity." Nature Structural & Molecular Biology 21, 423–425 (2014). dx.doi.org/10.1038/nsmb.2799
Journal information: Nature Structural & Molecular Biology