How knots can swap positions on a DNA strand
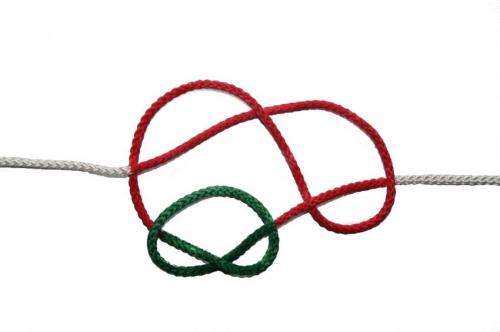

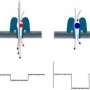
Physicists of Johannes Gutenberg University Mainz (JGU) and the Graduate School of Excellence "Materials Science in Mainz" (MAINZ) have been able with the aid of computer simulations to confirm and explain a mechanism by which two knots on a DNA strand can interchange their positions.
For this, one of the knots grows in size while the other diffuses along the contour of the former. Since there is only a small free energy barrier to swap, a significant number of crossing events have been observed in molecular dynamics simulations, i.e., there is a high probability of such interchange of positions.
"We assume that this swapping of positions on a DNA strand may also happen in living organisms," explained Dr. Peter Virnau of the JGU Institute of Physics, who performed the computer simulation together with his colleagues Benjamin Trefz and Jonathan Siebert.
The scientists expect that the mechanism may play an important role in future technologies such as nanopore sequencing, where long DNA strands are sequenced by being pulled though pores. Long DNA strands of more than 100,000 base pairs have an increasing chance of knots, which is relevant for sequencing.
More information: Benjamin Trefz, Jonathan Siebert, Peter Virnau, How molecular knots can pass through each other , Proceedings of the National Academy of Sciences, 19 May 2014 . DOI: 10.1073/pnas.1319376111
Journal information: Proceedings of the National Academy of Sciences
Provided by Universitaet Mainz