Chemists develop new technique for improving stomach cancer surgery results

When Stanford surgeon George Poultsides removes a tumor in the stomach or intestines, he takes out what he thinks is the entire mass and sends it to the pathology lab to evaluate. Completely removing each and every cancer cell is the best hope those patients have for a full recovery.
As much as half an hour later he finds out whether any cancerous cells remain at the edge of the tissue he removed. If yes, he takes out more tissue, sends it to pathology and waits for the answer.
Not only does this process extend the surgery, the information is wrong up to 30 percent of the time. Five days after the surgery, when the definitive test is complete, Poultsides knows whether he got all of the cancerous cells or whether he might need to call a patient back for another surgery.
This is where things stood when he got an email from chemistry postdoctoral fellow Livia Eberlin. She and her advisor, Professor Richard Zare, had an unusual idea for how chemical analysis could improve the odds of detecting cancer cells during surgery and prevent patients from needing to return. The results of that collaboration were published online Feb. 3 in the Proceedings of the National Academy of Sciences.
Learning new tricks
Eberlin chose to contact Poultsides from a website listing of cancer surgeons at Stanford Hospital because he appeared young and had a publication list that indicated he participated in research. "The unique thing at Stanford is proximity and Dr. Zare encouraged me to take advantage of that," said Eberlin, the lead author of the paper. "I had a feeling that younger clinicians would be more open to new technologies and decided to contact George."
Eberlin's expertise is in mass spectrometry, a tool not commonly used in a hospital setting. It takes a sample in one end, turns the molecules into charged particles, then detects how long it takes each charged molecule in that sample to migrate down a vacuum tube. The result is a jagged mountain range of tens of thousands of peaks, each representing a single chemical in the sample. The height of the peak indicates how much of that chemical the sample contained.
The idea was that maybe some of those peaks would be different in tissue samples that had cancerous cells versus those that didn't. If it worked, this mass spectrometry approach would be considerably faster than the hour-long screen pathologists do now, and more accurate.
Eberlin approached Poultsides about trying her idea in pancreatic cancer because she'd heard how hard those tumors are to treat, but Poultsides steered her toward stomach cancers. "I thought perhaps gastric cancer is where we have the biggest need," said Poultsides, a co-senior author of the paper and a member of the Stanford Cancer Institute. "Of all the cancers I work on, this is the one where we have the most frustration."
One holdup in developing the project was finding someone to fund it. It's not every day that chemists join teams of surgeons, pathologists and statisticians to solve problems in cancer treatment; the odd-couple partnership was a risk. Eventually they got a Stanford Hospital & Clinics Cancer Innovation Fund award, which supports unusual approaches to improving cancer treatment. "It was brave of them to fund this project," said Zare, a professor of chemistry and co-senior author of the paper. Zare is an affiliated member of Stanford's interdisciplinary Bio-X program and from that knew that unusual partnerships like this one can be unusually productive.
Poultsides and Eberlin made use of Stanford's tumor bank, which stores tissue donated by patients for research. They took some tissue known to have cancerous cells and some that didn't, and ran it all through the mass spectrometer. They ended up with roughly 10,000 peaks to analyze from each of about 500 sections of 62 frozen tissue samples.
Then, Zare called his colleague statistician Robert Tibshirani to help make sense of the results.
Finding what's important
Tibshirani, a professor of health research and policy and of statistics, has a long history of pulling useful information out of massive data sets like this one. "Most peaks aren't informative," he said. "You want to throw away what isn't helping you and pull out what's important."
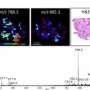
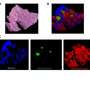
Tibshirani, who is also a co-author of the paper, pulled out the important peaks using a statistical technique called Lasso that he pioneered with Stanford colleagues almost 20 years ago. At the end of his analysis, about 120 peaks turned out to be important in distinguishing between samples that contain cancerous cells and those that don't. Their system analyzes those 120 peaks for each of the roughly 500 pixels of a tissue sample, and produces an image of the pixels color coded as cancerous or not.
The technique worked so well on banked tissue that they decided to go head-to-head against the current approach. When Poultsides and collaborator Jeffrey Norton, who is chief of surgical oncology, removed a tumor, they sent samples to the pathology lab to carry out the standard test and also called Eberlin. She ran across the street from the Zare lab with some ice and brought the sample back into the chemistry building to analyze.
A few days later, Eberlin sat down with collaborating pathologists Teri Longacre, Gerald Berry and David Bingham to compare the results from their technique with pathological results of the surgery samples. What they found is that the new technique was right almost every time.
The group members say they want to test the technique in a larger pool of stomach cancers to make sure it is as accurate as it seems, and also start working with other cancers in which it's not always clear whether the surgeon got the entire tumor. They also say the peaks that distinguish between cancer and normal cells could help point scientists to better understand what goes wrong in cancerous cells.
More information: Molecular assessment of surgical-resection margins of gastric cancer by mass-spectrometric imaging, www.pnas.org/cgi/doi/10.1073/pnas.1400274111
Journal information: Proceedings of the National Academy of Sciences
Provided by Stanford University