Suicidal bacteria: Biologists study unicellular organisms that occasionally poison themselves with a toxin

The cyanobacterium Synechocystis produces toxins that often lead to its own demise. The biologists Stefan Kopfmann and Prof. Dr. Wolfgang Hess from the University of Freiburg have determined the logic governing this mechanism.. Their findings have been published in the renowned periodicals Journal of Biological Chemistry (JBC) and Public Library of Science (PLoS ONE).
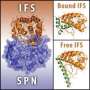
The cyanobacterium Synechocystis produces several toxins. However, most of the time they cannot become active because the unicellular organism usually only produces them together with an antitoxin that neutralizes their poisonous effect. This is a trick of nature: The genes for the toxin and the antitoxin are located together on a plasmid, i.e. a fragment of DNA that exists independently of the actual bacterial chromosome. In contrast to the toxin, the antitoxin is not very stable. When a cell loses the plasmid during cell division, both of the genes are lost. Since the toxin is more stable than the antitoxin and is thus effective for a longer period of time, these cells eventually die off. Hence, the toxin-antitoxin pairs constitute a natural selection mechanism that sees to it that only cells which retain the plasmid survive.
The plasmid pSYSA of the cyanobacterium Synechocystis has not one but seven different systems of this kind and is thus well protected. The reason for this is because in addition to the genes for the seven toxin-antitoxin pairs, the plasmid pSYSA possesses the genetic information for a bacterial immune system. If the plasmid with this system gets lost in cell division, several toxins thus see to it that the bacterium is killed. The fact that the genes responsible for it are combined with a high amount of toxin-antitoxin pairs indicates that this system has special significance for the cyanobacterial cell.
More information: Kopfmann, S. and Hess, W. Toxin antitoxin systems on the large defense plasmid pSYSA of Synechocystis sp. PCC 6803, Journal of Biological Chemistry. First Published on January 15, 2013, doi: 10.1074/jbc.M112.434100 jbc.M112.434100
Scholtz, I. et al. (2013) CRISPR-Cas Systems in the Cyanobacterium Synechocystis sp. PCC6803 Exhibit Distinct Processing Pathways Involving at Least Two Cas6 and a Cmr2 Protein. PLoS ONE 8(2): e56470. doi:10.1371/journal.pone.0056470
Journal information: Journal of Biological Chemistry , PLoS ONE
Provided by Albert Ludwigs University of Freiburg