Study suggests expanding the genetic alphabet may be easier than previously thought
A new study led by scientists at The Scripps Research Institute suggests that the replication process for DNA—the genetic instructions for living organisms that is composed of four bases (C, G, A and T)—is more open to unnatural letters than had previously been thought. An expanded "DNA alphabet" could carry more information than natural DNA, potentially coding for a much wider range of molecules and enabling a variety of powerful applications, from precise molecular probes and nanomachines to useful new life forms.
The new study, which appears in the June 3, 2012 issue of Nature Chemical Biology, solves the mystery of how a previously identified pair of artificial DNA bases can go through the DNA replication process almost as efficiently as the four natural bases.
"We now know that the efficient replication of our unnatural base pair isn't a fluke, and also that the replication process is more flexible than had been assumed," said Floyd E. Romesberg, associate professor at Scripps Research, principal developer of the new DNA bases, and a senior author of the new study. The Romesberg laboratory collaborated on the new study with the laboratory of co-senior author Andreas Marx at the University of Konstanz in Germany, and the laboratory of Tammy J. Dwyer at the University of San Diego.
Adding to the DNA Alphabet
Romesberg and his lab have been trying to find a way to extend the DNA alphabet since the late 1990s. In 2008, they developed the efficiently replicating bases NaM and 5SICS, which come together as a complementary base pair within the DNA helix, much as, in normal DNA, the base adenine (A) pairs with thymine (T), and cytosine (C) pairs with guanine (G).
The following year, Romesberg and colleagues showed that NaM and 5SICS could be efficiently transcribed into RNA in the lab dish. But these bases' success in mimicking the functionality of natural bases was a bit mysterious. They had been found simply by screening thousands of synthetic nucleotide-like molecules for the ones that were replicated most efficiently. And it had been clear immediately that their chemical structures lack the ability to form the hydrogen bonds that join natural base pairs in DNA. Such bonds had been thought to be an absolute requirement for successful DNA replication —a process in which a large enzyme, DNA polymerase, moves along a single, unwrapped DNA strand and stitches together the opposing strand, one complementary base at a time.
An early structural study of a very similar base pair in double-helix DNA added to Romesberg's concerns. The data strongly suggested that NaM and 5SICS do not even approximate the edge-to-edge geometry of natural base pairs—termed the Watson-Crick geometry, after the co-discoverers of the DNA double-helix. Instead, they join in a looser, overlapping, "intercalated" fashion. "Their pairing resembles a 'mispair,' such as two identical bases together, which normally wouldn't be recognized as a valid base pair by the DNA polymerase," said Denis Malyshev, a graduate student in Romesberg's lab who was lead author along with Karin Betz of Marx's lab.
Yet in test after test, the NaM-5SICS pair was efficiently replicable. "We wondered whether we were somehow tricking the DNA polymerase into recognizing it," said Romesberg. "I didn't want to pursue the development of applications until we had a clearer picture of what was going on during replication."
Edge to Edge
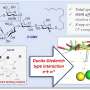
To get that clearer picture, Romesberg and his lab turned to Dwyer's and Marx's laboratories, which have expertise in finding the atomic structures of DNA in complex with DNA polymerase. Their structural data showed plainly that the NaM-5SICS pair maintain an abnormal, intercalated structure within double-helix DNA—but remarkably adopt the normal, edge-to-edge, "Watson-Crick" positioning when gripped by the polymerase during the crucial moments of DNA replication.
"The DNA polymerase apparently induces this unnatural base pair to form a structure that's virtually indistinguishable from that of a natural base pair," said Malyshev.
NaM and 5SICS, lacking hydrogen bonds, are held together in the DNA double-helix by "hydrophobic" forces, which cause certain molecular structures (like those found in oil) to be repelled by water molecules, and thus to cling together in a watery medium. "It's very possible that these hydrophobic forces have characteristics that enable the flexibility and thus the replicability of the NaM-5SICS base pair," said Romesberg. "Certainly if their aberrant structure in the double helix were held together by more rigid covalent bonds, they wouldn't have been able to pop into the correct structure during DNA replication."
An Arbitrary Choice?
The finding suggests that NaM-5SICS and potentially other, hydrophobically bound base pairs could some day be used to extend the DNA alphabet. It also hints that Evolution's choice of the existing four-letter DNA alphabet—on this planet—may have been somewhat arbitrary. "It seems that life could have been based on many other genetic systems," said Romesberg.
He and his laboratory colleagues are now trying to optimize the basic functionality of NaM and 5SICS, and to show that these new bases can work alongside natural bases in the DNA of a living cell.
"If we can get this new base pair to replicate with high efficiency and fidelity in vivo, we'll have a semi-synthetic organism," Romesberg said. "The things that one could do with that are pretty mind blowing."
More information: "KlenTaq polymerase replicates unnatural base pairs by inducing a Watson-Crick geometry," Nature Chemical Biology, 2012.
Journal information: Nature Chemical Biology
Provided by The Scripps Research Institute