Soil bacteria and pathogens share antibiotic resistance genes

(PhysOrg.com) -- Disease-causing bacteria’s efforts to resist antibiotics may get help from their distant bacterial relatives that live in the soil, new research at Washington University School of Medicine suggests.
The researchers found identical genes for antibiotic resistance in soil bacteria and in pathogens from clinics around the world. The matches prove that the two groups of bacteria have recently shared these genes but do not establish the direction of the sharing.
The results will be presented Feb. 20 at the annual meeting of the American Association for the Advancement of Science in Vancouver, British Columbia. The presentation is part of a panel discussion titled “Winning: Superbugs or Surveillance and Science?”
“A majority of the antibiotics used today are produced by soil bacteria, so it’s no surprise that the same bacteria have genes for resisting antibiotics,” says presenter Gautam Dantas, PhD, assistant professor of pathology and immunology. “Antibiotic resistance genes have likely been around for billions of years in the soil, but we wanted to take a first look at whether any of them are being exchanged with bacteria that cause human disease.”
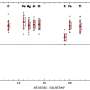
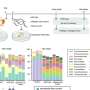
Using soil samples from sites in the United States, Dantas and his colleagues identified a series of antibiotic resistance genes in soil bacteria that can resist five classes of antibiotics, including forms of penicillin, sulfonamide and tetracycline. They found seven genes, of which several appear to be clustered together, that collectively employ all the known strategies for resisting antibiotics: blocking or ejecting the antibiotic from infected host cells, directly attacking the antibiotic or modifying the bacterial protein targeted by the antibiotic.
The same antibiotic resistance genes were present, often in similarly clustered arrangements, in samples of disease-causing bacteria from medical clinics around the world. Many of the matched genes were identical not only in the sections of the genes that code for proteins but also in nearby non-coding sections.
Bacterial DNA normally accumulates mutations and other alterations much more quickly than the DNA of humans. The lack of changes in the antibiotic resistance genes identified in the study suggests that the transfers of the genes must have occurred fairly recently in evolutionary history.
“We don’t yet know how much of a challenge these gene transfers are for our efforts to control infectious diseases,” says Kevin Forsberg, a graduate student in Dantas’ lab who led the research. “Are there a whole lot of these antibiotic resistance clusters being passed around, or did we just get lucky in discovering this potent group in our first assessment?”
“I suspect the soil is not a teeming reservoir of antibiotic resistance genes,” Dantas says. “But when we dump antibiotics into the environment, as our society does in a variety of contexts, we may be enriching that reservoir, and that may make antibiotic resistance genes more accessible to infectious bacteria.”
Dantas’ presentation will also feature earlier research he conducted on the presence of antibiotic resistance genes in human gut microbes.