Sewage treatment plants may contribute to antibiotic resistance problem
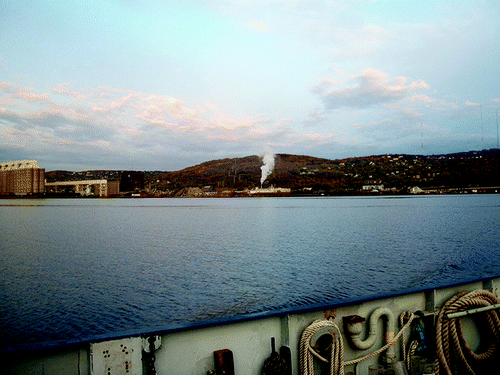
Water discharged into lakes and rivers from municipal sewage treatment plants may contain significant concentrations of the genes that make bacteria antibiotic-resistant. That's the conclusion of a new study on a sewage treatment plant on Lake Superior in the Duluth, Minn., harbor that appears in ACS' journal Environmental Science & Technology.
Timothy M. LaPara and colleagues explain that antibiotic-resistant bacteria — a major problem in medicine today — are abundant in the sewage that enters municipal wastewater treatment plants. Treatment is intended to kill the bacteria, and it removes many of the bacterial genes that cause antibiotic resistance. However, genes or bacteria may be released in effluent from the plant. In an effort to determine the importance of municipal sewage treatment plants as sources of antibiotic resistance genes, the scientists studied releases of those genes at the Duluth facility.
Although the Duluth facility uses some of the most advanced technology for cleaning wastewater — so-called tertiary treatment — the study identified it as an important source of antibiotic resistance genes. Sampling of water at 13 locations detected three genes, for instance, that make bacteria resistant to the tetracycline group of antibiotics, which are used to treat conditions ranging from acne to sexually transmitted diseases to anthrax and bubonic plague. LaPara's team says their research demonstrates that even the most high-tech sewage treatment plants may be significant sources of antibiotic resistance genes in waterways.
More information: Tertiary-Treated Municipal Wastewater is a Significant Point Source of Antibiotic Resistance Genes into Duluth-Superior Harbor, Environ. Sci. Technol., 2011, 45 (22), pp 9543–9549. DOI: 10.1021/es202775r
Abstract
In this study, the impact of tertiary-treated municipal wastewater on the quantity of several antibiotic resistance determinants in Duluth-Superior Harbor was investigated by collecting surface water and sediment samples from 13 locations in Duluth-Superior Harbor, the St. Louis River, and Lake Superior. Quantitative PCR (qPCR) was used to target three different genes encoding resistance to tetracycline (tet(A), tet(X), and tet(W)), the gene encoding the integrase of class 1 integrons (intI1), and total bacterial abundance (16S rRNA genes) as well as total and human fecal contamination levels (16S rRNA genes specific to the genus Bacteroides). The quantities of tet(A), tet(X), tet(W), intI1, total Bacteroides, and human-specific Bacteroides were typically 20-fold higher in the tertiary-treated wastewater than in nearby surface water samples. In contrast, the quantities of these genes in the St. Louis River and Lake Superior were typically below detection. Analysis of sequences of tet(W) gene fragments from four different samples collected throughout the study site supported the conclusion that tertiary-treated municipal wastewater is a point source of resistance genes into Duluth-Superior Harbor. This study demonstrates that the discharge of exceptionally treated municipal wastewater can have a statistically significant effect on the quantities of antibiotic resistance genes in otherwise pristine surface waters.
Provided by American Chemical Society