Paper shows enzyme-controlled movement of DNA polymer through a nanopore
Research published this week in Nature Nanotechnology shows a new method of enzyme-controlled movement of a single strand of DNA through a protein nanopore. The paper, by researchers at the University of California Santa Cruz (UCSC), represents a key step towards nanopore sequencing of DNA strands.
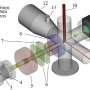
The publication describes the observation of single stranded DNA (ssDNA) as it translocates through a protein nanopore, alpha hemolysin (AHL). Movement of the ssDNA was controlled by polymerase-facilitated replication of individual DNA molecules. This movement could be initiated under electronic control. Polymerase activity was shown to be blocked in solution when the ssDNA was not at the nanopore opening, however capture of the strand by the pore removes a blocking strand of nucleotides and allows the polymerase to function on the trapped strand.
UCSC researchers are collaborating with Oxford Nanopore Technologies Ltd in the development of a new generation of electronic, single-molecule DNA sequencing technology. In the 'strand sequencing' method, current through a nanopore is measured as a DNA polymer passes through that pore. Changes in this current are used to identify the DNA bases on the DNA molecule, in sequence. This paper addresses a key challenge for DNA strand sequencing: fine control of the translocation of the DNA strand through the nanopore, at a rate that is consistent and slow enough to enable accurate identification of individual DNA bases. The Nature Nanotechnology work shows for the first time that the motion of a strand can be controlled using electronic feedback and that an enzyme can move a strand against a field while located on top of the nanopore.
"The techniques described in this paper are an advance towards electronic, single molecule DNA sequencing of DNA strands" said investigator Professor Mark Akeson of the University of California, Santa Cruz. "Electronic control of DNA translocation through a protein nanopore is a scientific goal that we have strived towards for years and these methods are now forming the basis for further work in our laboratories. We are excited by our collaboration with Oxford Nanopore, whose parallel nanopore sensing strategy is impressive."
More information: Replication of individual DNA molecules under electronic control using a protein nanopore. Felix Olasagasti, Kate R. Lieberman, Seico Benner, Gerald M. Cherf, Joseph M. Dahl, David W. Deamer and Mark Akeson Nature Nanotechnology September 2010. DOI: 10.1038/NNANO.2010.177
Provided by Feinstein Kean Healthcare