A fingerprint for genes
Cells may not have a mouth, but they still need to ingest substances from the external environment. If this process - known as endocytosis - is affected, it can lead to infectious diseases or cardio-vascular diseases, cancer, Huntington’s and diabetes. In cooperation with the Center for Information Services and High Performance Computing (ZIH) at the Dresden University of Technology, scientists from the Max Planck Institute of Molecular Cell Biology and Genetics therefore applied a new strategy to identify and characterize genes involved in endocytosis.
For that a combination of high-resolution microscopy and quantitative image analysis enabled the scientists to investigate the effects of a large number of genes. From their findings the scientists also hope to derive significant information about how infections could be prevented and diseases treated in future. (Nature, February 28, 2010, advance online publication)
Cells take up material from the outside by pinching off from their cell membrane vesicles that transport substances to different cellular organelles. Depending on what they contain, these vesicles and organelles - also known as endosomes - are transported to different locations within the cell, where their content is either re-distributed or broken down to recycle the basic building blocks. Endosomes are organised in a complex transport network that ensures a correct transport of a wide variety of substances towards their proper intracellular destination. However, the exact details of how, for instance, signalling molecules arrive at their respective destinations and transmit their information from the cell membrane to the nucleus are still largely unknown.
The new investigation strategy provided the scientists with previously unimagined insights into the highly complicated processes that take place in the cell. They discovered in their images, for example, that a failure of certain genes cause the arrest of vesicles in the cell periphery rather than being transported to the centre of the cell. Furthermore, different substances such as nutrients and growth factors are apparently guided to their destination by different set of genes and endocytosis is controlled by various signalling pathways. At the same time, the cells use endocytosis to carefully adjust the quantity of signal molecules on the cell membrane and in the endosomes - endocytosis and signalling pathways thus influence each other. All in all, more than 4,000 genes are directly or indirectly involved in endocytosis. "Our findings demonstrate that cells don't simply go out and ingest just any substances, handling them in the same manner. On the contrary, they have a very precise definition of what they need when and in what quantity, and also where it needs to get to in the cell," says Marino Zerial, Director at the Max Planck Institute of Molecular Cell Biology and Genetics.
Impaired endocytosis can cause disease
The enormous number of genes involved also reflects the significance of endocytosis for the cell and the entire organism. For example, signalling of important metabolic regulators like insulin depend on endocytosis but viruses and other pathogens also use endosomes to infect cells. Immune cells, for their part, swallow pathogens and digest them inside endosomes. Therefore, it might one day be possible to prevent dangerous infections if the endocytosis of viruses and bacteria could be selectively inhibited or if the pathogens could be destroyed more effectively in the endosomes. Even diseases like Alzheimer's and Huntington's are associated with a damaged transport system in the cells, as this prevents survival signals for nerve cells. Consequently, if the role of the different genes in endocytosis were known, it would in future be easier to develop potential treatments for these diseases.
Thanks to the new technology developed in Dresden, this will now be significantly easier. The Max Planck scientists succeeded in measuring the role of genes in endocytosis within a cell with unprecedented accuracy. To achieve this, they conducted quantitative analyses of microscopic images on the basis of dozens of different parameters and assigned each gene a certain function in the process. "We scanned and mapped the activity of all genes of the human genome. As a result, we were able to compile a quantitative profile of each gene - giving each gene an individual fingerprint, so to speak. This provides us with a more comprehensive understanding of how the various processes in the cell are integrated. This approach can now be applied to many different cellular mechanisms. We're well on the way to creating a 'virtual cell' by understanding all of the processes and different interactions", explains Marino Zerial.
New therapies call for new investigation strategies
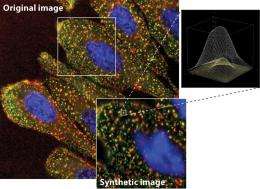
This was made possible by a combination of numerous technologies. The Dresden-based scientists blocked each of the approximately 23,000 expressed human genes one by one with the help of small interfering RNA molecules (siRNAs) that silence the respective gene. Next they used fluorescent dyes to mark two proteins which the cells under investigation took up in their endosomes - this made them visible to automated high-resolution microscopy and image-analysis software. Instead of analysing the microscopic images subjectively and on the basis of qualitative criteria, the scientists determined 62 different parameters for the cells and all visible endosomes in the images (multi-parametric image analysis). These criteria helped them to ascertain how blocking a gene affects endocytosis. They were therefore able to observe under the microscope how substance uptake in the cells changed if one of the human genes was inactivated.
"We developed a brand new strategy that combines numerous components into one big analysis system: a genomic RNAi screen, automated, high-resolution microscopy, multi-parametric image analysis - and computing power," reports Zerial. In the end, they had accumulated over two-and-a-half million images that needed to be analysed. This staggering volume of data could only be managed with a supercomputer, which was possible by the team's cooperation with the Center for Information Services and High Performance Computing (ZIH) at TU Dresden. A computer would have taken more than four million hours just to analyse the images to produce statistically reliable data. "But the incredible tour de force did more than bring significant new findings. It was also necessary as a means of providing a starting point for using this technology more efficiently and inexpensively in the future," says Zerial.
The next step will be to test the successful screening programme on primary cells or on model organisms simulating human disease conditions. This is when its true potential for drug development will be revealed. Ivan C. Baines, Director of Services and Facilities at the Max Planck Institute of Molecular Cell Biology and Genetics, outlines the future of the project: "If we can distinguish drug toxicity as a separate activity from drug efficacy, as is suggested by the quantitative profiling approach of this study, then we may be able to discover better drugs with fewer side effects."
More information: Systems survey of endocytosis by multiparametric image analysis, Nature, 28 February 2010, Advance Online Publication (AOP); doi:10.1038/nature08779
Provided by Max-Planck-Gesellschaft