Atomic force microscope will investigate muscle movement at molecular level
In a concrete bunker set upon bedrock on the Northern Arizona University campus lies an instrument delicate enough to manipulate a single molecule.
The atomic force microscope in the basement of the new Science and Health Building will be used to test a hypothesis developed by Kiisa Nishikawa, Regents' Professor of Biological Sciences, which could shift the foundation of our understanding of muscle movement.
Samrat Dutta, a post-doctoral scholar on Nishikawa's team, has already been putting in time on the AFM. He's relying on nine years of previous experience with a similar instrument while working on development of anti-cancer drugs.
"We are dealing with molecules in the nanometer range," said Dutta, who was recruited to help Nishikawa's team test the winding filament hypothesis. "We want to connect how change in the behavior of a single molecule in muscle contributes to overall behavior of how the muscle works."
The transdisciplinary research team received $1 million in funding from the W. M. Keck Foundation in 2014 specifically to pursue this work.
Nishikawa visualizes that a molecule called titin, the longest protein in humans, binds to a filament of actin (another protein) and winds around it to contribute to muscle contraction.

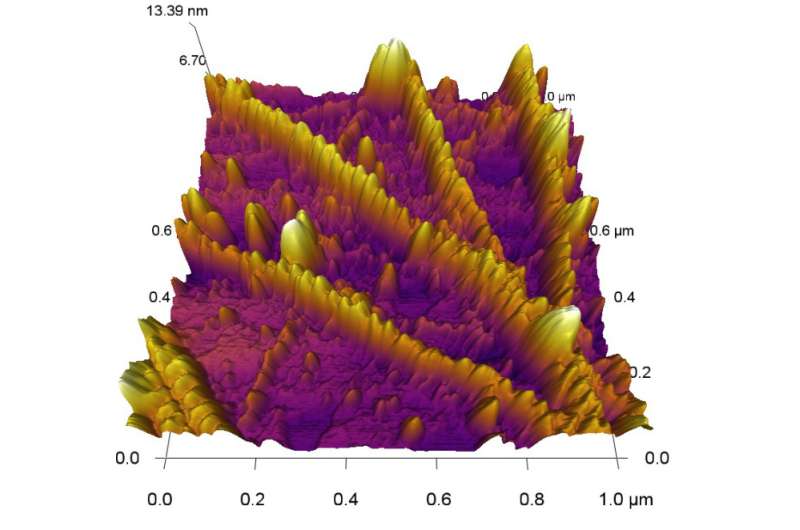
What the team hopes to see won't actually be a real-time image. Instead, two different techniques are used to produce something for researchers to see. In one, sophisticated software and statistical analysis are required to transform data generated by the microscope into a digital representation.
In the other, called force spectroscopy, researchers use a cantilever with a silicon tip that moves over the top of a substance, generating electrical signals. For this project, Dutta will attach a single molecule of titin to the tip, gently press it against a filament of actin, then measure the interaction.
"When we take off the cantilever, all the bonds involved in that interaction process will break," Dutta said. "We can measure those interactions and quantify how strongly a titin molecule binds to other proteins."
Attaching the titin molecule is achieved through surface chemistry, Dutta said. A covalent bond strongly binds a single molecule to the cantilever tip. How does he know it's only one?
"We have to play around with different concentration levels," Dutta said, pointing out that just getting the chemistry right is an active field of study in itself. That painstaking process contributes to atomic force microscopy being a slow technique.
"On a good day, I can get four or five images," Dutta said. "But the amount of information you get is huge."
Dutta's work so far has entailed polymerizing F-actin, a substrate to which actin can bind. As actin filaments—supplied by an outside lab—form on the surface, he images them.
"This is a dynamic process, so we want to test our experiment under physiological conditions," Dutta said.
Provided by Northern Arizona University