The beat goes on with a new model for artificial flagella

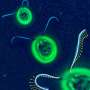
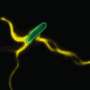
(Phys.org) —Eukaryotic flagella, whip-like organelles that elegantly propel microorganisms and pump fluid, seem to embody simplicity on the microscopic scale. But appearances can be deceptive: Flagella are composed of 650 different types of proteins.
Their jobs are vitally important. Flagella help sperm swim, sponges eat, and sweep mucus from the lungs, among other functions. Their length depends on their purpose but flagellas' structure and rhythmic, beating movement remain the same across functions and species.
That fluid movement is a highly sought-after capability in small-scale devices, such as microrobots. But scientists have struggled to build a simple, controllable model that can recreate it.
Now, Michael Hagan, associate professor of physics, and his lab, have built the first viable computer model to generate flagella-like movement with man-made structures.
Hagan's research was published online in the Journal of Royal Society Interface, coauthored by former Brandeis University postdoctoral fellows Raghunath Chelakkot and Arvind Gopinath, along with L. Mahadevan, professor of physics, biology, and applied mathematics, at Harvard University. The research was funded by the Material Research Science and Engineering Center (MRSEC) at Brandeis University.
Hagan's computer model significantly simplifies the motion of complex flagella, using spherical self-propelled particles called colloids in a structure resembling a string of beads. The colloids exert pressure on themselves, causing the filament to beat. To make the model viable, the team's first had to figure out the proper strength of the colloids' attachment. If the connections were too tight, the string would be stiff and unmoving; too loose, and it would be floppy and ineffective. Next, Hagan determined the length of the filament required for motion—too short or too long and it wouldn't be able to propel anything.
After determining the strength and length of the filament, the team anchored one end of it, as if to a cell wall, and observed those graceful, beating motions on their computer model.
"Because this system is so simple, and its construction so different from that of flagella, we should be able to elucidate the most fundamental features of flagella that give rise to and control motion. These features can be understood without having to unravel all 650 moving parts of a flagellum," Hagan says.
The research may pave the way to develop flagella-like microrobots to carry drugs to targeted cells or to create microfluidic devices that could pump and circulate fluid.
Provided by Brandeis University