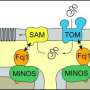
Study sheds new light on how cellular transport systems harness energy to perform their work inside the cell
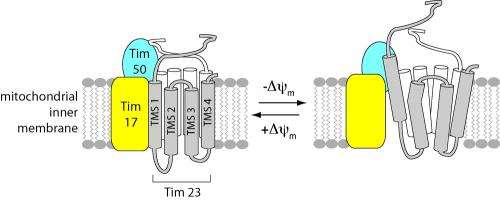
Using highly sensitive fluorescent probes, a team of scientists from the University of Connecticut has captured the never-before-seen structural dynamics of an important protein channel inside the cell's primary power plant – the mitochondrion.
The UConn team's study found that the channel complex - known as the translocase of the inner mitochondrial membrane 23 or TIM23 – is not only directly coupled to the energized state of the mitochondrial inner membrane as scientists have long suspected, it also changes its fundamental structure - altering the helical shape of protein segments that line the channel - when voltage along the membrane's electrical field drops.
The research, which appears this week in the peer-reviewed journal Nature Structural & Molecular Biology, explains how the energized state of the membrane drives the structural dynamics of membrane proteins and sheds new light on how cellular transport systems harness energy to perform their work inside the cell.
It also shows how fluorescent mapping at the subcelllar level may reveal new insights into the underlying causes of neurodegenerative and metabolic disorders associated with mitochondrial function.
In an overview of the research accompanying the paper's publication, Nikolaus Pfanner of the University of Freiberg in Germany and an international leader in the field of cellular protein trafficking, and several members of his research group, called the study "a major step towards a molecular understanding of a voltage-gated protein translocase."
"The molecular nature of voltage sensors in membrane proteins is a central question in biochemical research," Pfanner and his colleagues said. "The study…is not only of fundamental importance for our understanding of mitochondrial biogenesis, but also opens up new perspectives in the search for voltage-responsive elements in membrane proteins."
To conduct the study, UConn researchers incorporated cysteine residues modified with a fluorescent probe at specific positions along a transmembrane segment of a TIM23 complex derived from a common species of yeast, Saccharomyces cerevisiae. The team then monitored the probes in real time to observe how the channel's voltage-gating and structure responded to induced changes in the inner membrane's electrical field.
"It's an indirect way of looking at the structure of something, but because we are able to look into an actually functioning mitochondrion, it's given us a whole world of new information," says Nathan N. Alder, an assistant professor in the Department of Molecular and Cell Biology in UConn's College of Liberal Arts and Sciences and the research team's leader.
The study was supported by grants from the National Science Foundation, the National Institutes of Health and the Robert A. Welch Foundation.
"That the magnitude of the voltage gradient across the membrane could play a significant role in defining the structure of these proteins is probably one of the most significant elements of this research," Alder says.
The next phase of the research will look toward isolating the TIM23 protein channel complex in an artificial system to see if it continues to respond to voltage fluctuations outside of its natural habitat. The research team is also hoping to identify the particular parts of the protein complex that are acting as voltage sensors.
"Once we start to identify exactly what is the voltage sensor, we will have a better understanding of the translocase process and ultimately we can apply this knowledge to other kinds of protein transporters whose dysfunction has been implicated in the etiology of diseases such as cardiovascular disease and cancer," Alder says. "If their function is tied to the energized state of the membrane, we'll be able to see whether defects in that ability to couple to the membrane might be associated with the pathogenesis of these diseases."
More information: Structural changes in the mitochondrial Tim23 channel are coupled to the proton-motive force, Nature Structural & Molecular Biology, DOI: 10.1038/nsmb.2613
Journal information: Nature Structural & Molecular Biology
Provided by University of Connecticut