Protein production: going viral: Architecture of essential human transcription factor revealed
(Phys.org)—A research team of scientists from EMBL Grenoble and the IGBMC in Strasbourg, France, have, for the first time, described in molecular detail the architecture of the central scaffold of TFIID: the human protein complex essential for transcription from DNA to mRNA. The study, published today in Nature, opens new perspectives in the study of transcription and of the structure and mechanism of other large multi-protein assemblies involved in gene regulation.
By controlling the transcription of DNA into messenger RNA, TFIID forms the cornerstone of the machinery that controls gene expression in our cells. Despite its crucial role, very little was known about its architecture. TFIID is present at very low levels in cells, and it is a very large protein complex made of 20 subunits: this combination largely prevented previous attempts to purify it and decipher its structure and function in molecular detail. Even the most advanced methods for recombinant protein production met their limits when trying to produce its various subunits in the right proportions.
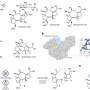
The solution to this bottleneck came from studying the strategy certain viruses, such as Coronaviruses, use when they replicate: they produce very long protein chains that are then divided into individual proteins. Mimicking this technique led to highly abundant and correctly assembled complexes of the core scaffold of TFIID (comprising 10 subunits), which could be purified and analysed at high resolution by combining electron microscopy and data from X-ray crystallography.
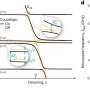
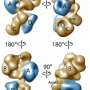
This ground-breaking analysis reveals the inner workings of the core-complex of human TFIID in unprecedented detail. It shows that some of its subunits adopt a very defined structure, whereas other parts appear to adopt intricate, extended geometries winding like worms through the complex, holding it together. The overall architecture of the complex is symmetric; however, the authors describe how it becomes asymmetric when it binds to other subunits to finally form the complete TFIID complex.
"We know now in some detail what the core of TFIID looks like, and what happens when further subunits are bound. We believe that we have opened the door to determining the architecture of the entire human TFIID complex in the near future, and likewise of other large multiprotein assemblies involved in gene regulation, and to explain their roles in catalysing biological function," concludes Imre Berger, coordinator of the study at EMBL.
More information: The architecture of human general transcription factor TFIID core complex. Christoph Bieniossek, Gabor Papai, Christiane Schaffitzel, Frederic Garzoni, Maxime Chaillet, Elisabeth Scheer, Petros Papadopoulos, Laszlo Tora, Patrick Schultz and Imre Berger. Published online in Nature on the 6 January 2013. dx.doi.org/10.1038/nature11791
Abstract
The initiation of gene transcription by RNA polymerase II is regulated by a plethora of proteins in human cells. The first general transcription factor to bind gene promoters is transcription factor IID (TFIID). TFIID triggers pre-initiation complex formation, functions as a coactivator by interacting with transcriptional activators and reads epigenetic marks1–3. TFIID is a megadalton-sized multiprotein complex composed of TATA-box-binding protein (TBP) and 13 TBP-associated factors (TAFs)3. Despite its crucial role, the detailed architecture and assembly mechanism of TFIID remain elusive. Histone fold domains are prevalent in TAFs, and histone-like tetramer and octamer structures have been proposed in TFIID4–6. A functional core-TFIID subcomplex was revealed in Drosophila nuclei, consisting of a subset of TAFs (TAF4, TAF5, TAF6, TAF9 and TAF12)7. These core subunits are thought to be present in two copies in holo-TFIID, in contrast to TBP and other TAFs that are present in a single copy8, conveying a transition from symmetry to asymmetry in the TFIID assembly pathway. Here we present the structure of human core-TFIID determined by cryoelectron microscopy at 11.6A ° resolution. Our structure reveals a two-fold symmetric, interlaced architecture, with pronounced protrusions, that accommodates all conserved structural features of the TAFs including the histone folds.Wefurther demonstrate that binding of one TAF8–TAF10 complex breaks the original symmetry of core-TFIID. We propose that the resulting asymmetric structure serves as a functional scaffold to nucleate holo-TFIID assembly, by accreting one copy each of the remaining TAFs and TBP.
Journal information: Nature
Provided by European Molecular Biology Laboratory