Research reveals new understanding of X chromosome inactivation
(Phys.org)—In a paper published in the Nov. 21 issue of Cell, a team led by Mauro Calabrese, a postdoctoral fellow at the University of North Carolina in the lab of Terry Magnuson, chair of the department of genetics and member of the UNC Lineberger Comprehensive Cancer Center, broadens the understanding of how cells regulate silencing of the X chromosome in a process known as X-inactivation.
"This is a classic example of a basic research discovery. X-inactivation is a flagship model for understanding how non-coding RNAs orchestrate large-scale control of gene expression. In the simplest terms, we are trying to understand how cells regulate expression of their genes. Our findings are relevant across the board—by understanding how normal cells function we can apply that knowledge to similar situations in the understanding and treatment of disease," said Calabrese.
Proper regulation of the X chromosome plays a crucial role in mammalian development. Females inherit a pair of X chromosomes from their parents, and the process of X-inactivation shuts down one of these two Xs.
"Males have XY. Females have two Xs. One of those Xs needs to get shut off. If it does not, it's not compatible with life. It's how we have evolved to equalize doses between males and females," said Calabrese.
While the manner in which the X chromosome is deactivated has been actively studied for 50 years, the exact mechanisms that regulate the process remain a mystery. Calabrese's research used high-throughput sequencing to determine the location and activity of chromosomes with far greater accuracy than previous research.
"Basically, this is using the sequencing technology as a high resolution microscope," said Calabrese.
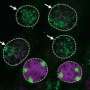
Under a microscope, the inactive X chromosome (Xi) appears as a cloud-like structure, because it is covered with a non-coding RNA known as Xist. In the traditional model of X-inactivation, genes located inside the cloud are completely silenced, with 15 percent of the genes from the inactive X chromosomes escaping to become active.
"The prevailing thought was that genes that escaped X inactivation were pulled out of the core and expressed out there," said Calabrese.
The work of Calabrese's team complicates the current model of X-inactivation by finding indications of gene activity inside the Xist cloud and the presence of inactive genes outside the cloud, both of which would not have been thought possible in the prevailing model.
"It's kind of a subtle thing, but mechanistically it is a big difference," said Calabrese.
Inside the Xist cloud, sequencing discovered traces of DNase I sensitivity, a feature usually linked to transcription activity. While other markers associated with transcription were absent, the presence of DNase I sensitivity suggested that the nucleus did recognize the inactive X as usable DNA, but an unknown suppressive mechanism was preventing genes from being activated.
"We were surprised to see that. If they were totally silent, you would expect this to be not there… This suggests that transcription factors or other proteins that bind DNA are still accessing the inactive X," said Calabrese.
The other surprising findings involve the 15 percent of "escaper" genes from the inactive X. Calabrese found evidence that active genes were found both inside and outside the Xist cloud, and that silenced genes that lay alongside active genes outside of the Xist cloud remained inactive.
"If X-inactivation was a strict nuclear barrier, then pulling a gene outside the barrier would turn it on, but it has got to be more than that because when an inactivated gene that is beside an escaper is outside this domain, it is still turned off," said Calabrese.
The presence of DNase I sensitivity within the Xist cloud and the finding of inactive genes outside of the cloud suggest that a site-specific mechanism is regulating genes on the chromosome in a more subtle way than the binary "on/off" function posited by the prevailing model. The exact mechanism for this remains unknown. Although Calabrese believes that Xist still plays a role, its exact function and whether other factors influence X-inactivation remain questions for future research.
"We know that Xist is required to turn off the inactive X. We know that. We have no idea how" said Calabrese.
Beyond revising the understanding of how X-inactivation works, Calabrese said that deeper understanding of the function of Xist could reveal more about the role of other non-coding RNAs in cellular development. These RNAs could become useful targets for future therapies and drug development.
"We know that too much expression of the wrong non-coding RNAs can lead to cancer. Also, forced expression of other non-coding RNAs can prevent cancer. Generally, we do not know how these RNAs work," said Calabrese.
Journal information: Cell
Provided by University of North Carolina Health Care