Kinetochore structure reveals how it takes hold
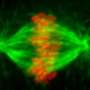
(Phys.org) -- With the first-ever three-dimensional image of an isolated kinetochore – the bulky molecular machine that connects a chromosome to the long, thin microtubules that tug it to one end of a dividing cell -- scientists can now see how the machine establishes and maintains its grip, even as the microtubule tip it holds onto shrinks away. Maintaining that grip is essential to ensure that new cells receive the appropriate allotment of chromosomes.
Tamir Gonen, a group leader at the Howard Hughes Medical Institute’s Janelia Farm Research Campus, led the effort to determine the kinetochore structure, in collaboration with Sue Biggins’ lab at the Fred Hutchinson Cancer Research Center in Seattle. His team’s report of the new structure was published online on August 12, 2012, in the journal Nature Structural and Molecular Biology.
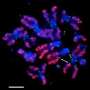
When cells divide, it is crucial that the resulting daughter cells receive the same genetic information contained in the parent – no more, no less. To enable this, cells prepare to divide by first making copies of their chromosomes. As the cell elongates, pairs of identical chromosomes separate and each partner is drawn toward an opposite end of the cell by a long, dynamic component of the cellular skeleton known as a microtubule. To form an attachment site for the microtubule, hundreds of proteins come together on the chromosome to assemble the kinetochore complex.
The kinetochore must maintain its grip on the microtubule even as it is being pulled toward the end of the elongating cell. That’s particularly impressive, Biggins says, because microtubules are dynamic, constantly growing and retracting at their ends. During cell division, microtubules diminish in length, bringing their tips nearer the cell’s edge. For a kinetochore, Biggins says, “it’s as if you’re climbing a rope and someone is constantly pulling the rope out from under you. The kinetochore somehow hangs on to this dynamic polymer, even as the polymer is disassembling right under it.”
Scientists had proposed a few strategies that might enable the kinetochore to stay connected to the vanishing tip of a microtubule. One model suggested that it might form a ring that floats freely along the length of the microtubule. Alternatively, some research had suggested, it might establish attachments to multiple binding sites along the microtubule, so that some of the attachments could come and go as the microtubule changed shape. But without being able to actually see the kinetochore, Biggins says, it was difficult to determine which model was right.
Even in yeast, in which the kinetochore is relatively simple, many copies of each of 38 different proteins must come together to form a functional kinetochore. “This thing is enormous,” Gonen says, noting that at 100 nanometers across, a kinetochore is four times the size of another complex cellular structure, the ribosome. Further, it is inherently dynamic. While subcomplexes have been reconstituted, no one has assembled an entire functional kinetochore.
With persistence, Bungo Akiyoshi, a student in Biggins’ lab, devised a method to purify the unwieldy structure. Gonen’s brother Shane, a technician in the Biggins lab, could then use electron microscopy to try to image the kinetochores as they grabbed on to microtubules. Still, the unstable complexes often began to degrade before the images were captured, resulting in pictures that of kinetochores in various states of disrepair. It was impossible to tell just by looking which of the shapes on the grid represented intact kinetochores. “From a structural biology perspective, it was a mess. We had particles that were varying in size from very small to very large, with variable shapes,” Tamir says. “And when we started we had no idea what we were looking for. There were a lot of things there, and we really couldn’t figure out what was what.”
“It was only after the Biggins lab started generating mutants that were missing various components of the kinetochore that things started making sense,” says Tamir, who was an HHMI Early Career Scientist at the University of Washington at the time. “Then we could figure out what particles were indeed kinetochores and assign the position of some components.”
Over two years, Shane collected terabytes of data and the researchers began putting the pieces of the puzzle together. With statistical analyses, they determined which features were reproducible and indicative of the structure of the fully assembled complex, and by the time Tamir moved his lab across the country to Janelia Farm in 2011, they had a pretty good idea of what the kinetochore looked like.
Still, they were missing critical data that would reveal what it looked like in three dimensions. Gonen proposed that his Janelia Farm lab use electron tomography to generate a three-dimensional image of the kinetochore. When Dan Shi, the Gonen lab’s electron microscopy specialist, arrived at Janelia Farm, he set out to collect the necessary data. Using the same grids that Shane had prepared to collect the original images, Shi focused on a few particles that appeared to be intact kinetochore complexes, then collected a series of images of each one from different angles. Matt Iadanza, a graduate student in the Gonen lab then merged the data to produce the three-dimensional structures. “Essentially what you have are all these side views of the particle, and you combine those to get a three-dimensional structure,” Gonen explains. When they did that, he says, “things started making even a little bit more sense. Instead of a flattened view of the top of the particle, we now had a density that we could rotate around and have a look.”
Gonen and Biggins say it was shocking how much their team’s structures reflected models that researchers had imagined several years earlier, based on biochemical data. “It was amazing to see it look like what people had pictured in their head,” Biggins says. “When you look at it, you can really imagine its behavior.” Their images showed that the kinetochore is shaped something like the palm of a hand. A large domain in the center might grab onto the chromosome, the researchers say, while the spokes that radiate out from that center form attachments to the microtubule.
Some, but not all, of the kinetochores in their images formed a ring structure, and Biggins says the structures they have observed suggest that both of the previously proposed models for microtubule binding are likely true: kinetochores can form the ring structure that biochemists had proposed might facilitate their attachment to microtubules, but that the ring is not necessary.
“These are very early days for this structure,” Gonen says, noting that the large size and dynamic nature of the kinetochore will be a challenge as structural biologists work toward a higher-resolution picture of the complex. “But with this unprecedented look, people can now begin to explain some of the biochemical data about how kinetochores control chromosome segregation.”
More information: Abstract: www.hhmi.org/research/groupleaders/gonen.html
Journal information: Nature Structural and Molecular Biology
Provided by Howard Hughes Medical Institute