The physics of going viral: Researchers measure the rate of DNA transfer from viruses to bacteria
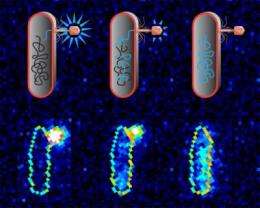
Researchers at the California Institute of Technology (Caltech) have been able, for the first time, to watch viruses infecting individual bacteria by transferring their DNA, and to measure the rate at which that transfer occurs. Shedding light on the early stages of infection by this type of virus—a bacteriophage—the scientists have determined that it is the cells targeted for infection, rather than the amount of genetic material within the viruses themselves, that dictate how quickly the bacteriophage's DNA is transferred.
"The beauty of our experiment is we were able to watch individual viruses infecting individual bacteria,"says Rob Phillips, the Fred and Nancy Morris Professor of Biophysics and Biology at Caltech and the principal investigator on the new study. "Other studies of the rate of infection have involved bulk measurements. With our methods, you can actually watch as a virus shoots out its DNA."
The new methods and results are described in a paper titled "A Single-Molecule Hershey–Chase Experiment," which will appear in the July 24 issue of the journal Current Biology and currently appears online. The lead authors of that paper, David Van Valen and David Wu, completed the work while graduate students in Phillips's group.
In the well-known 1952 Hershey-Chase experiment, Alfred Hershey and Martha Chase of the Carnegie Institution of Washington in Cold Spring Harbor convincingly confirmed earlier claims that DNA—and not protein—was the genetic material in cells. To prove this, the researchers used bacteriophages, which are able to infect bacteria using heads of tightly bundled DNA coated in a protein shell. Hershey and Chase radiolabeled sulfur, contained in the protein shell but not in the DNA, and phosphorous, found in the DNA but not in the protein shell. Then they let the bacteriophages infect the bacterial cells. When they isolated the cells and analyzed their
contents, they found that only the radioactive phosphorous had made its way into the bacteria, proving that DNA is indeed the genetic material. The results also showed that, unlike the viruses that infect humans, bacteriophages transmit only their genetic information into their bacterial targets, leaving their "bodies" behind.
"This led, right from the get-go, to people wondering about the mechanism—about how the DNA gets out of the virus and into the infected cell," Phillips says. Several hypotheses have focused on the fact that the DNA in the virus is under a tremendous amount of pressure. Indeed, previous work has shown that the genetic material is under more pressure within its protein shell than champagne experiences in a corked bottle. After all, as Phillips says, "There are 16 microns [16,000 nanometers] of DNA in a tiny 50-nanometer-sized shell. It's like taking 500 meters of cable from the Golden Gate Bridge and putting it in the back of a FedEx truck."
Phillips's group wanted to find out whether that pressure plays a dominant role in transferring the DNA. Instead, he says, "What we discovered is that the thing that mattered most was not the pressure in the bacteriophage, but how much DNA was in the bacterial cell."
The researchers used a fluorescent dye to stain the DNA of two mutants of a bacteriophage known as lambda bacteriophage—one with a short genome and one with a longer genome—while that DNA was still inside the phage. Using a fluorescence microscope, they traced the glowing dye to see when and over what time period the viral DNA transferred from each phage into an E. coli bacterium. The mean ejection time was about five minutes, though that time varied considerably.
This was markedly different from what the group had seen previously when they ran a similar experiment in a test tube. In that earlier setup, they had essentially tricked the bacteriophages into ejecting their DNA into solution—a task that the phages completed in less than 10 seconds. In that case, once the phage with the longer genome had released enough DNA to make what remained inside the phage equal in length to the shorter genome, the two phages ejected DNA at the same rate. Therefore, Phillips's team reasoned, it was the amount of DNA in the phage that determined how quickly the DNA was transferred.
But Phillips says, "What was true in the test tube is not true in the cell." E. coli cells contain roughly 3 million proteins within a box that is roughly one micron (1,000 nanometers) on each side. Less than 10 nanometers separate each protein from its neighbors. "There's no room for anything else," Phillips says. "These cells are really crowded."
And so, when the bacteriophages try to inject their DNA into the cells, the factor that limits the rate of transfer is how jam-packed those cells are. "In this case," Phillips says, "it had more to do with the recipient, and less to do with the pressure that had built up inside the phage."
Looking toward the future, Phillips is interested in using the methods he and his team have developed to study different types of bacteriophages. He also wants to investigate various molecules that could be helping to actively pull the viral DNA into the cells. In the case of a bacteriophage called T7, for instance, previous work has shown that the host cell actually grabs onto the DNA and begins making copies of its genes before the virus has even delivered all the DNA into the cell. "We're curious whether that kind of mechanism is in play with the lambda bacteriophage," Phillips says.
The current findings have implications for the larger question of how biomolecules like DNA and proteins cross membranes in general, and not just into bacteria. Cells are full of membranes that divide them into separate compartments and that separate entire cells from the rest of the world. Much of the business of cellular life involves getting molecules across those barriers. "This process that we've been studying is one of the most elementary examples of what you could call polymer translocation or getting macromolecules across membranes," Phillips says. "We are starting to figure out the physics behind that process."
Journal information: Current Biology
Provided by California Institute of Technology