Researchers provide atomic view of a histone chaperone
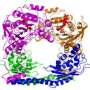
Mayo Clinic researchers have gained insights into the function of a member of a family of specialized proteins called histone chaperones. Using nuclear magnetic resonance (NMR) spectroscopy and X-ray crystallography, they have determined the 3-D structure and interactions of the histone chaperone Rtt106 down to the atomic details. The findings are published in the journal Nature.
"The interactions we described are important for gene silencing and the DNA damage response," says senior author Georges Mer, Ph.D., a Mayo Clinic structural biologist. "This is exciting because our newfound knowledge will help us better understand these fundamental cellular processes."
In cells, our DNA is part of a structure called chromatin, comprised of proteins, the majority called histones, which are wrapped with the DNA. Associated with the histones is another group of proteins called histone chaperones, which promote the proper assembly or disassembly of the chromatin during the times our DNA is replicated or repaired when damaged. Their dysfunction has been linked to cancer, aging and other diseases.
The Discovery
Before this Mayo study, scientists knew that the histone chaperone Rtt106 helped in the deposition of histones -- specifically, a complex of histones H3 and H4 -- onto the replicating DNA. They did not understand how Rtt106 does this, given that it does not possess any of the known requirements. Histone H3 is in a modified form where one of its amino acids, lysine 56, is acetylated. Rtt106 does not seem to have an acetylated lysine reader domain.
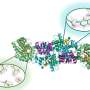
Mayo researchers discovered two novel domains in Rtt106 that take on this role. One, termed the homodimerization domain, allows two molecules of Rtt106 to be linked so they can cooperate in binding histones H3 and H4. The other, called the double PH domain, is responsible for sensing the acetylated lysine of H3 and further reinforces the interaction. The combined actions of the two domains of Rtt106 enable it to perform the chaperoning function efficiently and properly. This is the first time anyone has described this mode of specific association between a histone chaperone and a modified histone complex.
Provided by Mayo Clinic