Giant virus, tiny protein crystals show X-ray laser's power and potential

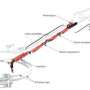
Two studies published in the February 3 issue of Nature demonstrate how the unique capabilities of the world's first hard X-ray free-electron laser -- the Linac Coherent Light Source, located at the Department of Energy's SLAC National Accelerator Laboratory -- could revolutionize the study of life.
In one study, an international research team used the LCLS to demonstrate a shortcut for determining the 3-D structures of proteins. The laser's brilliant pulses of X-ray light pulled structural data from tiny protein nanocrystals, avoiding the need to use large protein crystals that can be difficult or impossible to prepare. This could lop years off the structural analysis of some proteins and allow scientists to decipher tens of thousands of others that are out of reach today, including many involved in infectious disease.
In a separate paper, the same team reported making the first single-shot images of intact viruses, paving the way for snapshots and movies of molecules, viruses and live microbes in action.
Led by Henry Chapman of the Center for Free-Electron Laser Science at the German national laboratory DESY and Janos Hajdu of Sweden's Uppsala University, the team of more than 80 researchers from 21 institutions performed these experiments in December 2009, just two months after the LCLS opened for research. Their studies are the first to demonstrate the power and potential of the LCLS for biology.
"The LCLS beam is a billion times brighter than previous X-ray sources, and so intense it can cut through steel," Chapman said. "Yet these incredible X-ray bursts are used with surgical, microscopic precision and exquisite control, and this is opening whole new realms of scientific possibilities," including the ability to observe atoms moving and chemical bonds forming and breaking in real time.
Outrunning a laser blast
In the experiments, scientists sprayed viruses or nanocrystals into the path of the X-ray beam and zapped them with bursts of laser light. Each strobe-like laser pulse is so brief —a few millionths of a billionth of a second long—that it gathers all the information needed to make an image before the sample explodes.
Hajdu had proposed this method nearly a decade earlier. Researchers at Arizona State University, Lawrence Livermore National Laboratory, SLAC and Uppsala spent years developing specialized equipment for injecting samples into the beam, and Germany's Max Planck Advanced Study Group brought in a 10-ton, $7 million instrument called CAMP to record every single photon of data with a fast, ultra-sensitive X-ray camera for later analysis.
Tests at DESY and Lawrence Berkeley National Laboratory showed that the concept worked at lower X-ray energies. "But as you go to higher energies, can you still outrun the damage?" said team member Michael Bogan, a SLAC staff scientist and principal investigator at the PULSE Institute for Ultrafast Energy Science, jointly located at SLAC and Stanford University. The answer, he said, was yes: "The physics still holds."
A big payoff from tiny crystals
The protein structure experiments were led by Chapman and Arizona State's John Spence and Petra Fromme. They chose as their target Photosystem I, a biological factory in plant cells that converts sunlight to energy during photosynthesis. It's one of an important class of proteins known as membrane proteins that biologists and drug developers are eager to understand better.
Embedded in cell membranes, these proteins control traffic in and out of the cell and serve as docking points for infectious agents and disease-fighting drugs; in fact, they are the targets of more than 60 percent of the drugs on the market. Yet scientists know the structures of only six of the estimated 30,000 membrane proteins in the human body, given the difficulty of turning them into big crystals for conventional X-ray analysis.
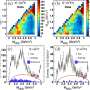
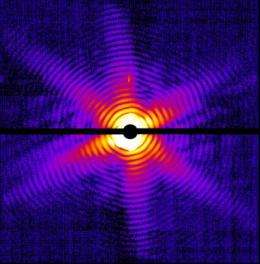
To get around this bottleneck, the researchers squirted millions of nanocrystals containing copies of Photosystem I across the X-ray beam. Laser pulses hit the crystals at various angles and scattered into the detector, forming the patterns needed to reconstitute images. Each crystal immediately vaporized, but by the time the next pulse arrived another crystal had moved into the bull's eye.
The team combined 10,000 of the three million snapshots they took to come up with a good match for the known molecular structure of Photosystem I.
"I attended several meetings this summer where this work was presented and I was extraordinarily excited by it," Michael Wiener of the University of Virginia, who was not involved in the research, said of the results. He leads one of nine institutes set up by the National Institutes of Health to decipher the structures of membrane proteins. "Preparation of these nanocrystals is likely to be very, very much easier than the larger crystals used to date," Wiener said, leaving scientists more time and money to find out how these important biomolecules work.
The team is scheduled to return to the LCLS this month to repeat the experiments with X-ray laser pulses that are much faster and deliver four times as much energy as they did in the initial round. If the physics still hold, future images should capture the extraordinarily complex structure of Photosystem I in atom-by-atom detail.
Portraits of a virus
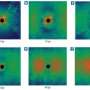
For the second experiment, the team went a step beyond nanocrystals to no crystals at all. Led by Hajdu, they made single-shot portraits of individual virus particles. These snapshots are a step toward eventually producing stop-action movies of chemical changes taking place in molecules and within living cells.
Biologists have long dreamed of making images of viruses, whole microbes and living cells without freezing, slicing or otherwise disturbing them. This is one of the goals of the LCLS, and the researchers tested its capabilities on Mimivirus, the world's largest known virus, which infects amoebas.
Of the hundreds of Mimiviruses hit by the LCLS beam, two produced enough data to allow scientists to reconstitute their images. The images show the 20-sided structure of the Mimi's outer coat and an area of denser material inside, which may represent its genetic material. Shorter, brighter pulses focused to a smaller area should greatly improve the resolution of these images to reveal details as small as one nanometer, the team wrote in their Feb. 3 Nature report.
Getting a detailed picture of the internal structure of an individual virus "would be a great achievement," said team member Jean-Michel Claverie, director of the Structural & Genomic Information Lab in Marseille and one of the scientists who discovered Mimi's viral nature.
"This is a brand-new way to look at a biological object," he said. "This will allow us to address not only the questions related to the internal structure of the virus, but its intrinsic variability from one individual virus particle to the next—a microscopic variability that might play a fundamental role in evolution."
The team returned to the LCLS in January to look at the Mimivirus at X-ray wavelengths that should maximize the amount of contrast and detail in the images. They will be analyzing the results in the months to come.
SLAC Director Persis Drell, who sat in a control room packed with scientists as raw data from the two experiments came in, said the experience was thrilling—and so is the potential for biology and medicine.
"This first data and these first papers are really just the first view of a new research frontier," she said. "They represent a turning point for the LCLS, demonstrating new technologies that will be great steps forward."
Provided by SLAC National Accelerator Laboratory