This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Unlocking the world of bacteria—researchers introduce new approach to make bacteria amenable to genetic engineering
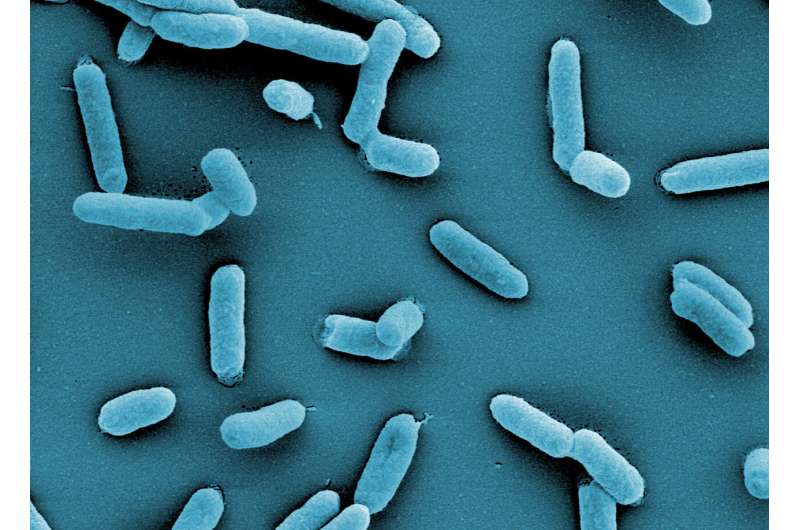
Bacteria populate virtually every habitat on Earth, including within and on our own bodies. Understanding and engineering bacteria can lead to new methods for diagnosing, treating, and preventing infections. Additionally, it presents opportunities to protect crops from disease and create sustainable cell factories for chemical production, reducing environmental impact—just a few of the many benefits to society.
To unlock these advantages, scientists need the ability to manipulate the genetic content of these bacteria. However, a longstanding bottleneck in genetically engineering bacteria has been the efficient transformation of DNA, the process of introducing foreign DNA into a cell. This has limited its application to only a small subset of microbes.
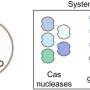
A major obstacle is the presence of restriction-modification systems. These protective systems mark the bacterial genome with a unique methylation pattern and destroy incoming foreign DNA lacking this pattern.
Overcoming this barrier requires adding the bacterium's pattern to the DNA, a process that is strain-specific and involves multiple DNA methyltransferases. These enzymes attach methyl groups, small chemical groups containing one carbon atom bonded to three hydrogen atoms, to DNA bases. Current methods to replicate or circumvent these DNA methylation patterns are labor-intensive and not easily scalable, necessitating new approaches.
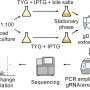
Addressing this challenge, a team led by the Helmholtz Institute for RNA-based Infection Research (HIRI), a site of the Braunschweig Helmholtz Centre for Infection Research (HZI) in cooperation with the Julius-Maximilians-Universität Würzburg (JMU), has introduced a novel approach to recreate such patterns and enhance DNA transformation. They called it IMPRINT, which stands for Imitating Methylation Patterns Rapidly IN TXTL.
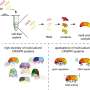
As part of this method, the researchers use a cell-free transcription-translation (TXTL) system—a liquid mixture that can produce ribonucleic acids (RNAs) and proteins from added DNA—to express a bacterium's specific set of DNA methyltransferases. The enzymes are then used to methylate DNA prior to its delivery into the target bacterium.
A wholly new application
"IMPRINT represents an entirely new use of TXTL. While TXTL is widely employed for various purposes, including producing hard-to-express proteins or as affordable diagnostic tools, it has not previously been utilized to overcome barriers to DNA transformation in bacteria," says Chase Beisel, head of the RNA Synthetic Biology department at the HIRI and professor at the JMU Medical Faculty. He spearheaded the study in collaboration with researchers from North Carolina State University (NC State) in Raleigh, U.S. Their findings were published today in the journal Molecular Cell.
Compared to existing methods, IMPRINT offers speed and simplicity. "Current approaches require either laboriously purifying individual DNA methyltransferases or expressing them in E. coli, which often proves cytotoxic," says Justin M. Vento, first author of the study who completed the work as a Ph.D. student in the Department of Chemical and Biomolecular Engineering at NC State. "These methods can take days to weeks and only reconstitute a fraction of the bacterium's methylation pattern."
The researchers demonstrated that IMPRINT could express a diverse array of DNA methyltransferases. Furthermore, these enzymes could be combined to recreate complex methylation patterns. This greatly enhanced DNA transformation in bacteria such as the pathogen Salmonella and the probiotic Bifidobacteria, including a challenging-to-transform strain of the latter, less-studied bacterium.
The basis for new antibiotics and cell-based therapies
The potential applications in modern medicine and research are extensive: IMPRINT can improve DNA transformation in clinical isolates of bacterial pathogens and in bacteria that combat infections, such as commensal bacteria or those producing antibacterial compounds. Genetic modification of these microbes could lead to new classes of antibiotics and cell-based therapies.
The research team aims to expand the use of IMPRINT. "We want to make a wide variety of bacterial pathogens genetically tractable for research," Beisel says. He hopes that IMPRINT will be widely adopted by the research community.
"Until now, certain bacteria have been favored as models simply because they are easier to genetically manipulate. We are hopeful that, by using IMPRINT, researchers will be able to focus on the most important bacterial strains, such as those with increased virulence or antibiotic resistance," Beisel concludes.
More information: Justin M. Vento et al, A cell-free transcription-translation pipeline for recreating methylation patterns boosts DNA transformation in bacteria, Molecular Cell (2024). DOI: 10.1016/j.molcel.2024.06.003
Journal information: Molecular Cell
Provided by Helmholtz Association of German Research Centres