This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers create new AI pipeline for identifying molecular interactions

Understanding how proteins interact with each other is crucial for developing new treatments and understanding diseases. Thanks to computational advances, a team of researchers led by Assistant Professor of Chemistry Alberto Perez have developed an algorithm to identify these molecular interactions.
Perez's research team included two graduate students from UF, Arup Mondal and Bhumika Singh, and a handful of researchers from Rutgers University and Rensselaer Polytechnic Institute. The team published their findings in Angewandte Chemie International Edition.
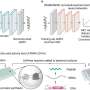
Named the AF-CBA Pipeline, this innovative tool offers unparalleled accuracy and speed in pinpointing the strongest peptide binders to a specific protein. It does this by using AI to simulate molecular interactions, sorting through thousands of candidate molecules to identify the molecule that interacts best with the protein of interest.
The AI-driven approach allows the pipeline to perform these actions in a fraction of the time it would take humans or traditional physics based-approaches to accomplish the same task.
"Think of it like a grocery store," Perez explained. "When you want to buy the best possible fruit, you have to compare sizes and aspects. There are too many fruits to try them all of course, so you compare a few before making a selection. This AI method, however, can not only try them all, but can also reliably pick out the best one."
Typically, the proteins of interest are the ones that cause the most damage to our bodies when they misbehave. By finding what molecules interact with these problematic proteins, the pipeline opens avenues for targeted therapies to combat ailments such as inflammation, immune dysregulation, and cancer.
"Knowing the structure of the strongest peptide binder in turn helps us in the rational designing of new drug therapeutics," Perez said.
The groundbreaking nature of the pipeline is enhanced by its foundation on pre-existing technology: a program called AlphaFold. Developed by Google Deepmind, AlphaFold uses deep learning to predict protein structures. This reliance on familiar technology will be a boon for the pipeline's accessibility to researchers and will help ensure its future adoption.
Moving forward, Perez and his team aim to expand their pipeline to gain further biological insights and inhibit disease agents. They have two viruses in their sights: murine leukemia virus and Kaposi's sarcoma virus. Both viruses can cause serious health issues, especially tumors, and interact with as-of-now unknown proteins.
"We want to design novel libraries of peptides," Perez said. "AF-CBA will allow us to identify those designed peptides that bind stronger than the viral peptides."
More information: Arup Mondal et al, A Computational Pipeline for Accurate Prioritization of Protein‐Protein Binding Candidates in High‐Throughput Protein Libraries, Angewandte Chemie International Edition (2024). DOI: 10.1002/anie.202405767
Journal information: Angewandte Chemie International Edition
Provided by University of Florida