This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
Unveiling the genetic tapestry of tree growth: A model for Populus euphratica development
A comprehensive understanding of the genetic architecture of tree growth, a complex interplay of genetics between the plant's above- and below-ground parts, remains undefined in plant studies. Research has increasingly focused on understanding how genes regulate growth, employing advanced methods such as transcriptome analysis and genome-wide association studies. Yet, these studies often narrow their focus to a few genes or traits, overlooking the holistic view of organ integrity and the full genetic network.
Therefore, there is a need for a comprehensive gene model approach, leveraging high-throughput genotyping technologies, to fully grasp the genetic mechanisms shaping complex plant phenotypes and their development.
In January 2024, Plant Phenomics published a research article titled "Genome-Wide Network Analysis of Above- and Below-Ground Co-growth in Populus euphratica."
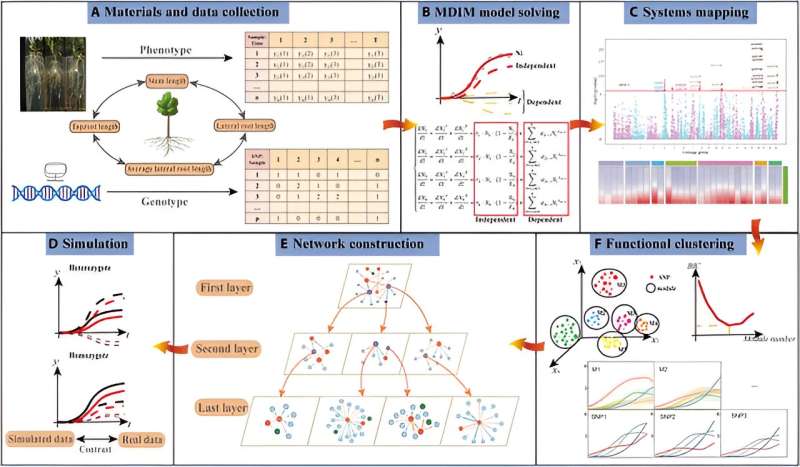
In this study, researchers introduce a pioneering computational model designed to uncover the genetic underpinnings of tree growth in Populus euphratica, focusing on the developmental dynamics and interactions between above- and below-ground traits. By integrating advanced methodologies like systems mapping, functional clustering, and evolutionary game theory, the model successfully delineates the genetic contributions and network topology driving phenotypic formation.
Specifically, the research explores the interactions among four critical traits—stem length, taproot length, lateral root length, and average lateral root length—using a multi-dimensional interactive model (MDIM). This approach not only fits the growth curves of these traits with high accuracy but also illuminates the complex interplay among them, revealing distinct time-varying growth characteristics and the influence of genetic interactions.
The findings indicate that the taproot exhibits superior growth compared to other traits, especially in early development stages, likely catering to the plant's need for water and nutrients. Furthermore, the analysis of significant quantitative trait loci (QTLs) identifies 54 key SNPs across the genome, contributing to the understanding of the genetic regulation of these traits. Notably, the model's ability to simulate the growth and genetic effects of Populus euphratica under various conditions validated its reliability and accuracy.
In addition, the construction of a multilayer large-scale interactive network, drawing inspiration from ecosystem structures, enables a holistic view of the genome-wide genetic architecture. This network, comprising multiple modules with differing genetic effects, showcases the complexity of genetic interactions in growth traits. Key QTLs are identified and their biological functions annotated, shining a light on their roles in plant development. The network's hierarchical structure, ranging from global gene interactions to specific SNP effects, provides unprecedented insights into the genetic basis of complex traits.
Overall, this study not only advances the genetic understanding of tree growth but also sets a precedent for the analysis of complex traits across different species. The model's flexibility and scalability suggest its applicability to a wide range of biological questions, highlighting its potential to revolutionize our approach to genetic research and plant breeding.
More information: Kaiyan Lu et al, Genome-Wide Network Analysis of Above- and Below-Ground Co-growth in Populus euphratica, Plant Phenomics (2023). DOI: 10.34133/plantphenomics.0131
Provided by TranSpread





















