This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers discover molecular 'barcode' used by bacteria to secrete toxins
Researchers at McMaster University have discovered a molecular "barcode" system used by disease-causing bacteria to distinguish between beneficial and toxic molecules.
Published in the Proceedings of the National Academy of Sciences (PNAS), the new study shows that many bacteria can figuratively scan genetic codes to learn which proteins to keep and which proteins to expel into the environment.
According to researchers, those proteins that are expelled are often toxic to human cells, making the ability to differentiate between proteins vital to a bacterium's capacity for causing infectious disease.
"Proteins are one of the fundamental building blocks of life," explains John Whitney, an associate professor in the Department of Biochemistry and Biomedical Sciences at McMaster and the lead investigator on the study. "They quite literally allow bacterial pathogens to do everything that they do. And while the vast majority of proteins remain inside bacteria to carry out functions like metabolism, there is a very small subset that act outside of the organism—like toxins."
Whitney, a member of the Michael G. DeGroote Institute for Infectious Disease Research, says that although the bacterial secretion system and the toxins themselves have long been studied, it was not understood how bacteria discriminated between toxic and non-toxic proteins prior to the study.
Whitney's lab, led by biochemistry graduate students Prakhar Shah and Timothy Klein (now a postdoctoral fellow at the University of California, San Francisco), questioned how three fundamentally different toxins could all be secreted by the same bacterial secretion system.
"There were no known similarities between the toxins—they don't look anything alike, and they don't do anything similar," Whitney says. "Our rationale was, for them to all pass through the same protein secretion machine, there must be something common between them."
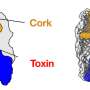
Sure enough, there was. Each toxin shared a "domain," which Whitney colloquially compares to a barcode. He says that while the barcode was shared by all three toxins under study, it was absent from the other 3,000 or so proteins in the bacteria, indicating that it serves as the export signal.
Shah, who co-first authored the study with Klein, says the team was able to prove this concept experimentally through a combination of genetic, biochemical, and structural approaches, including important "X-ray crystallography studies" that allowed them to purify the proteins and get a clearer view of the so-called barcodes and how they functioned.
The research team believes that this new information could eventually have a range of important biotechnology and infectious disease-related applications. In particular, Shah notes that the findings are relevant to our understanding of an array of gram-positive pathogens, including the kinds of bacteria responsible for serious infectious diseases like tuberculosis and listeriosis.
"A lot of pathogens use this system," Shah says. "Therefore, our discovery has important implications on our understanding of the virulence strategies used by a wide range of human pathogens."
More information: Structure of a tripartite protein complex that targets toxins to the type VII secretion system, Proceedings of the National Academy of Sciences (2024). DOI: 10.1073/pnas.2312455121. doi.org/10.1073/pnas.2312455121
Journal information: Proceedings of the National Academy of Sciences
Provided by McMaster University