This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Using single-antibodies as a new tool to build bio-circuitry
By using single-antibodies, Professor Hirohide Saito (Department of Life Science Frontiers) and his team of researchers, Shodai Komatsu and Assistant Professor Hirohisa Ohno, have developed a novel system to control gene expression in response to any target molecule inside cells, and they have employed it to design various synthetic biological circuits, including one for cell-specific genome editing.
The paper is published in the journal Nature Communications.
For cells to function properly and adapt to changes in the internal and external environment, there is a constant need to sense changes and react to them appropriately. For instance, cells have evolved a wide range of transcriptional (i.e., DNA→RNA), translational (i.e., RNA→protein), and even post-translational (i.e., protein modifications such as phosphorylation, or adding a phosphate group to specific locations on a protein) regulatory mechanisms to detect changes in nutrient and metabolite levels to modulate nutrient uptake, metabolism, and waste metabolite removal properly.
Another example is the immune cells in our bodies that constantly try to sense foreign DNA or RNA as signs of invading organisms and turn on certain genes to initiate an appropriate defensive response to potential threats.
In synthetic biology, scientists have leveraged natural transcriptional and translational regulators to customize biological circuits to control cellular functions in desirable ways. As we enter a new era of bioengineering and regenerative medicine, the demand for novel ways to build bio-circuitry is greater than ever.
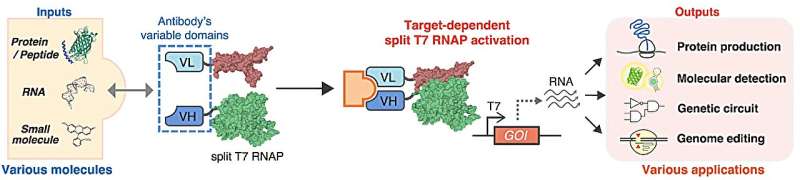
In their recent study, Saito and his team took a page out of nature's instruction manual and ingeniously applied antibodies, highly evolved proteins capable of binding a broad range of targets, such as DNA, RNA, proteins, and essentially any small molecules, to detect specific molecules as inputs and control transgene transcription as outputs.
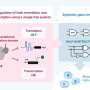
Specifically, antibodies from most mammals contain a variable region, a three-dimensional structure formed by a heavy and light chain, that binds a certain substrate (target) with high affinity. The researchers recognized that when the target molecule is present inside cells, it will interact separately with the heavy and light chains, effectively bringing them into proximity.
They therefore attached the variable region (heavy and light chains separately) from antibodies known to interact with specific target molecules to two parts of a split T7 RNA polymerase (RNAP), a protein designed to spontaneously reassemble when brought close together and transcribe RNA from an artificial DNA transgene, to drive expression of reporter genes as an output to signal positive target molecule detection.
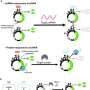
In the study, the researchers revealed the power and flexibility of their so-called target-dependent RNAP (TdRNAP) system to sense a variety of target molecules, including an antigenic peptide from the yeast transcription factor GCN4, the FLAG peptide, enhanced green fluorescence protein (EGFP), a specific RNA sequence from the hepatitis C virus (HCV), and the fluorescent small molecule, fluorescein, by simply switching between different variable region sequences for target detection.
In addition, by combining different RNAP variants into the system, they also demonstrated the creation of multilayer biological circuits for either signal amplification or orthogonal signal transduction. Aside from using this TdRNAP system simply for target molecule detection, the research team also demonstrated its application for cell-specific genome editing by using the system to induce guide RNA (gRNA) expression to trigger the CRISPR/Cas9-mediated deletion of a transgene previously incorporated into the genome of an experimental cell line.
The research team believes their new TdRNAP system to be an invaluable new tool for constructing bio-circuity to detect a variety of intracellular molecules and control cell functions. This system holds promise for potential applications in improving the efficacy and safety of future gene and cell therapies.
More information: Shodai Komatsu et al, Target-dependent RNA polymerase as universal platform for gene expression control in response to intracellular molecules, Nature Communications (2023). DOI: 10.1038/s41467-023-42802-5
Journal information: Nature Communications
Provided by Kyoto University