This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Finding a balance in antibiotic medicine: Can we kill bad bacteria while preserving the good?
A new study led by scientists at University of St Andrews has found that antibiotics used to treat tuberculosis kill other potentially useful bacteria.
The work has been published in The Lancet Microbe.
Human life and microorganisms evolved together mostly in a mutually beneficial relationship. Billions of these microorganisms live happily inside the body while others live outside on the skin. Together they are called microbiota. The good microbiota help strengthen our immunity against diseases, as well as helping to turn the food we eat into nutrients which nourish our bodies. A tiny minority of microbiota cause disease, which compels us to use medicines known as antibiotics that kill them.
However, while these antibiotics are effective in killing the disease-causing microbiota, studies have shown that they also kill the good ones.
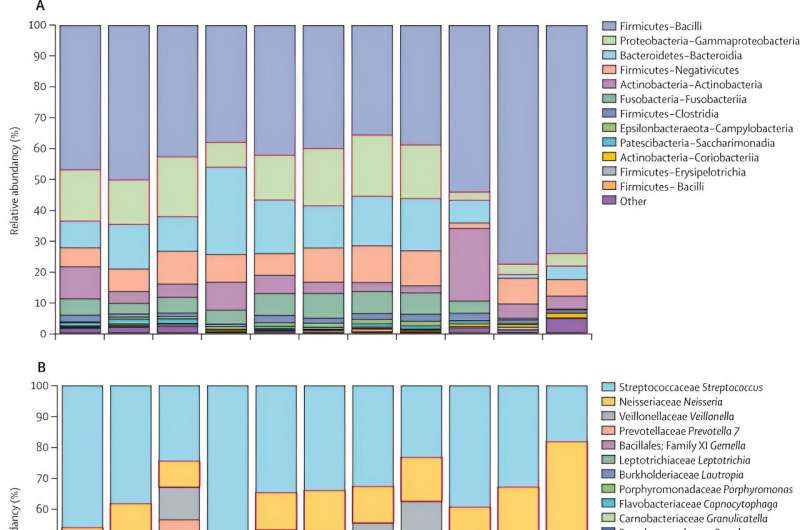
The "killing effect" was more pronounced in the first two weeks of the patient taking antibiotics, after which the bacteria began to recover, achieving a similar level of abundancy as before treatment within two months. Tuberculosis is often treated with combinations of four antibiotics. Seven of these combinations were studied.
While all combinations had a depressing effect on the microbial community, only two combinations—one containing 35 milligrams per kilogram of rifampicin, and the other containing 20 milligrams per kilogram of rifampicin, supplemented by 400mg of moxifloxacin—achieved significant killing of the microbiota.
Importantly, microbiota recovered faster in the latter combination which contained a lower dose of rifampicin. This implies that a carefully balanced antibiotic course can achieve the purpose of killing the disease-causing bacteria while preserving the useful ones that the body needs to recover to full health.
Commenting on these findings, lead researcher Dr. Wilber Sabiiti (School of Medicine) said, "For a long time, assessment of drug safety has only focused on the effect caused against human body organs such as the liver and the brain. This has neglected the effect caused against the useful microorganisms that live with humans and make them thrive. Many studies prior to ours have shown how the presence of these microorganisms helps educate human immunity to resist infections and prevent allergies and diseases like asthma. It is therefore crucial that the view of drug safety is expanded to include the safety of useful microorganisms in the human body."
It is notable that for this study, researchers used RNA, a genetic molecule that is closely associated with live cells. This enabled them to measure only microorganisms that were alive before and after exposure to antibiotics. Most previous studies of microbiota have used DNA, which is a stable genetic molecule that survives for many months and years after cell death. Therefore, measuring DNA of bacteria especially after exposure to antibiotics does not indicate whether or not the measurement is of live bacteria.
Despite being under antibiotic pressure, the microbiota recovered within two months of treatment in all combinations of antibiotics except one. Future studies will need to investigate whether this recovery is due to replenishment from dietary sources, or to bacteria acquiring antibiotic-resistant genes.
It is well known that long exposure to antibiotics makes bacteria devise ways to survive by developing or acquiring genes that enable them resist killing by antibiotics. To stop the lung and gut, which contain millions of microorganisms, from becoming "brewing factories" of antibiotic-resistant superbugs, studies must be done to understand how bacteria in these organs are responding to antibiotic pressure and devise means to stop emergence of antibiotic resistance.
This research is the result of collaboration between the study participants and researchers from Tanzania, and researchers from universities of Radboud in Netherlands, Munich in Germany, and St Andrews in the UK on behalf of the Pan African Consortium for Evaluation of anti-TB Antibiotics (PanACEA).
More information: Effect of seven anti-tuberculosis treatment regimens on sputum microbiome: a retrospective analysis of the HIGHRIF study 2 and PanACEA MAMS-TB clinical trials, The Lancet Microbe (2023). DOI: 10.1016/S2666-5247(23)00191-X , www.thelancet.com/journals/lan … (23)00191-X/fulltext
Journal information: The Lancet Microbe
Provided by University of St Andrews